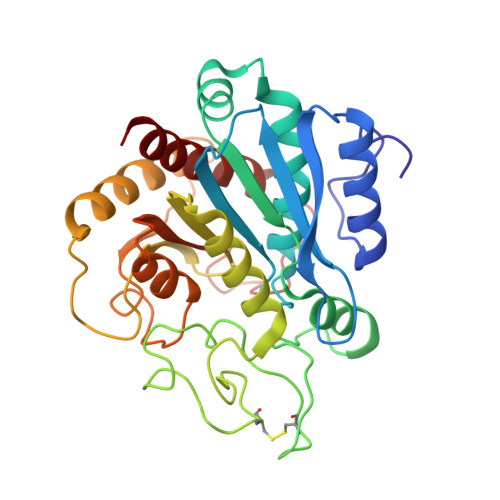
Crystal structure of the complex of carboxypeptidase A with a strongly bound phosphonate in a new crystalline form: comparison with structures of other complexes.
Kim, H., Lipscomb, W.N.(1990) Biochemistry 29: 5546-5555
- PubMed: 2386784
- DOI: https://doi.org/10.1021/bi00475a019
- Primary Citation of Related Structures:
6CPA - PubMed Abstract:
O-[[(1R)-[[N-(Phenylmethoxycarbonyl)-L-alanyl]amino]ethyl] hydroxyphosphinyl]-L-3-phenyllacetate [ZAAP(O)F], an analogue of (benzyloxycarbonyl)-Ala-Ala-Phe or (benzyloxycarbonyl)-Ala-Ala-phenyllactate, binds to carboxypeptidase A with great affinity (Ki = 3 pM). Similar phosphonates have been shown to be transition-state analogues of the CPA-catalyzed hydrolysis [Hanson, J. E., Kaplan, A. P., & Bartlett, P. A. (1989) Biochemistry 28, 6294-6305]. In the present study, the structure of the complex of this phosphonate with carboxypeptidase A has been determined by X-ray crystallography to a resolution of 2.0 A. The complex crystallizes in the space group P2(1)2(1)2(1) with cell dimensions a = 61.9 A, b = 67.2 A, and c = 76.2 A. The structure of the complex was solved by molecular replacement. Refinement of the structure against 20,776 unique reflections between 10.0 and 2.0 A yields a crystallographic residual of 0.193, including 140 water molecules. The two phosphinyl oxygens of the inhibitor bind to the active-site zinc at 2.2 A on the electrophilic (Arg-127) side and 3.1 A on the nucleophilic (Glu-270) side. Various features of the binding mode of this phosphonate inhibitor are consistent with the hypothesis that carboxypeptidase A catalyzed hydrolysis proceeds through a general-base mechanism in which the carbonyl carbon of the substrate is attacked by Zn-hydroxyl (or Zn-water). An unexpected feature of the bound inhibitor, the cis carbamoyl ester bond at the benzyloxycarbonyl linkage to alanine, allows the benzyloxycarbonyl phenyl ring of the inhibitor to interact favorably with Tyr-198. This complex structure is compared with previous structures of carboxypeptidase A, including the complexes with the potato inhibitor, a hydrated keto methylene substrate analogue, and a phosphonamidate inhibitor. Comparisons are also made with the complexes of thermolysin with some phosphonamidate inhibitors.
- Gibbs Chemical Laboratory, Harvard University, Cambridge, Massachusetts 02138.
Organizational Affiliation: