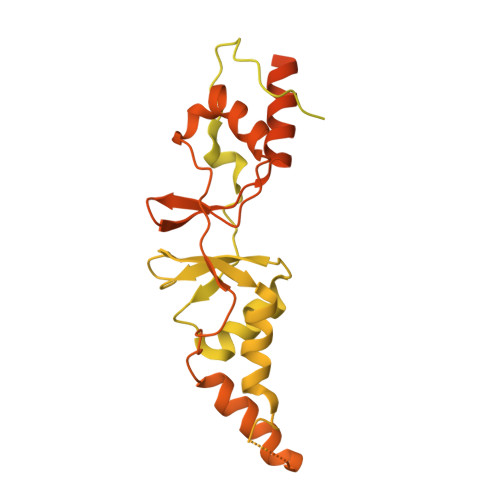
Structure of the CLC-1 chloride channel fromHomo sapiens.
Park, E., MacKinnon, R.(2018) Elife 7
- PubMed: 29809153 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.36629
- Primary Citation Related Structures:
6COY, 6COZ - PubMed Abstract:
CLC channels mediate passive Cl - conduction, while CLC transporters mediate active Cl - transport coupled to H + transport in the opposite direction. The distinction between CLC-0/1/2 channels and CLC transporters seems undetectable by amino acid sequence. To understand why they are different functionally we determined the structure of the human CLC-1 channel. Its 'glutamate gate' residue, known to mediate proton transfer in CLC transporters, adopts a location in the structure that appears to preclude it from its transport function. Furthermore, smaller side chains produce a wider pore near the intracellular surface, potentially reducing a kinetic barrier for Cl - conduction. When the corresponding residues are mutated in a transporter, it is converted to a channel. Finally, Cl - at key sites in the pore appear to interact with reduced affinity compared to transporters. Thus, subtle differences in glutamate gate conformation, internal pore diameter and Cl - affinity distinguish CLC channels and transporters.
- Laboratory of Molecular Neurobiology and Biophysics, Howard Hughes Medical Institute, The Rockefeller University, New York, United States.
Organizational Affiliation: