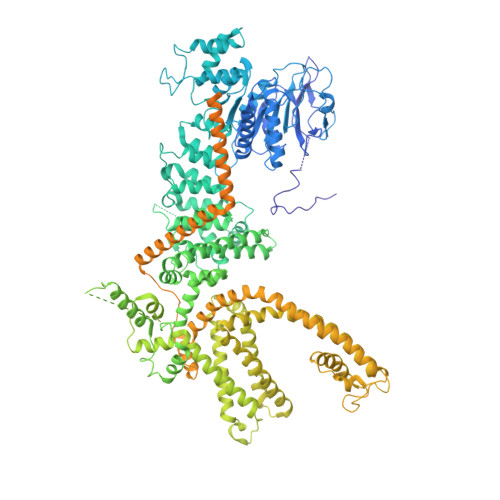
Structure of a TRPM2 channel in complex with Ca2+explains unique gating regulation.
Zhang, Z., Toth, B., Szollosi, A., Chen, J., Csanady, L.(2018) Elife 7
- PubMed: 29745897 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.36409
- Primary Citation Related Structures:
6CO7 - PubMed Abstract:
Transient receptor potential melastatin 2 (TRPM2) is a Ca 2+ -permeable cation channel required for immune cell activation, insulin secretion, and body heat control. TRPM2 is activated by cytosolic Ca 2+ , phosphatidyl-inositol-4,5-bisphosphate and ADP ribose. Here, we present the ~3 Å resolution electron cryo-microscopic structure of TRPM2 from Nematostella vectensis , 63% similar in sequence to human TRPM2, in the Ca 2+ -bound closed state. Compared to other TRPM channels, TRPM2 exhibits unique structural features that correlate with its function. The pore is larger and more negatively charged, consistent with its high Ca 2+ selectivity and larger conductance. The intracellular Ca 2+ binding sites are connected to the pore and cytosol, explaining the unusual dependence of TRPM2 activity on intra- and extracellular Ca 2+ . In addition, the absence of a post-filter motif is likely the cause of the rapid inactivation of human TRPM2. Together, our cryo-EM and electrophysiology studies provide a molecular understanding of the unique gating mechanism of TRPM2.
- Laboratory of Membrane Biophysics and Biology, The Rockefeller University, New York, United States.
Organizational Affiliation: