Structural Mechanism of Functional Modulation by Gene Splicing in NMDA Receptors.
Regan, M.C., Grant, T., McDaniel, M.J., Karakas, E., Zhang, J., Traynelis, S.F., Grigorieff, N., Furukawa, H.(2018) Neuron 98: 521-529.e3
- PubMed: 29656875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.neuron.2018.03.034
- Primary Citation Related Structures:
6CNA - PubMed Abstract:
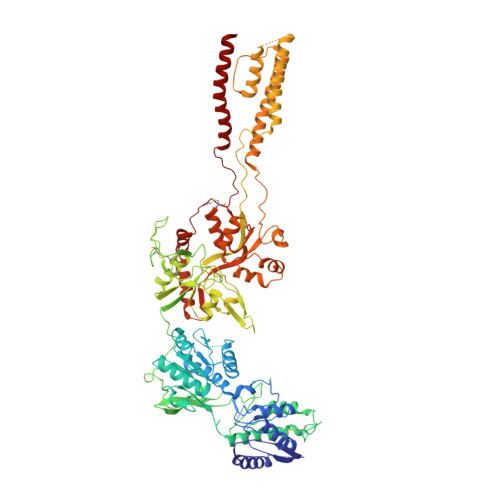
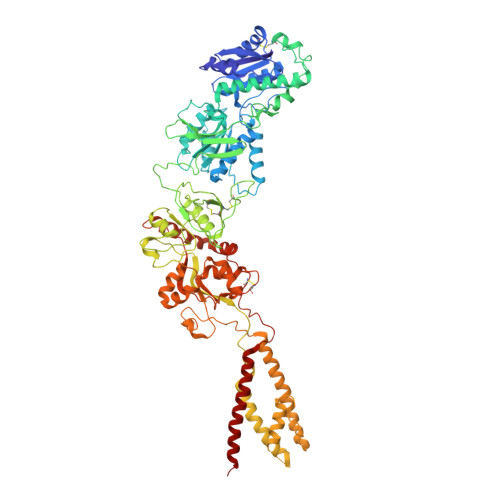
Alternative gene splicing gives rise to N-methyl-D-aspartate (NMDA) receptor ion channels with defined functional properties and unique contributions to calcium signaling in a given chemical environment in the mammalian brain. Splice variants possessing the exon-5-encoded motif at the amino-terminal domain (ATD) of the GluN1 subunit are known to display robustly altered deactivation rates and pH sensitivity, but the underlying mechanism for this functional modification is largely unknown. Here, we show through cryoelectron microscopy (cryo-EM) that the presence of the exon 5 motif in GluN1 alters the local architecture of heterotetrameric GluN1-GluN2 NMDA receptors and creates contacts with the ligand-binding domains (LBDs) of the GluN1 and GluN2 subunits, which are absent in NMDA receptors lacking the exon 5 motif. The unique interactions established by the exon 5 motif are essential to the stability of the ATD/LBD and LBD/LBD interfaces that are critically involved in controlling proton sensitivity and deactivation.
- WM Keck Structural Biology Laboratory, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY 11724, USA.
Organizational Affiliation: