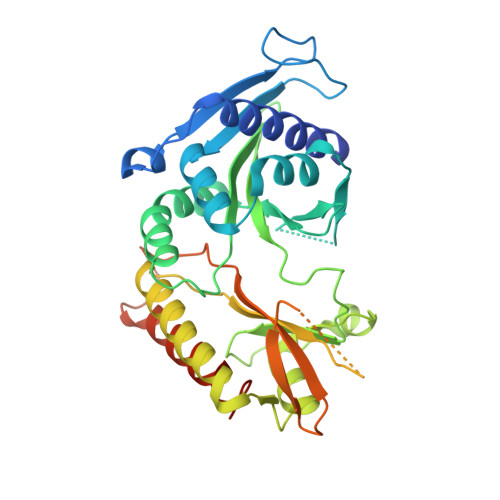
Structural insights into lethal contractural syndrome type 3 (LCCS3) caused by a missense mutation of PIP5K gamma.
Zeng, X., Uyar, A., Sui, D., Donyapour, N., Wu, D., Dickson, A., Hu, J.(2018) Biochem J 475: 2257-2269
- PubMed: 29959184 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20180326
- Primary Citation Related Structures:
6CMW, 6CN2, 6CN3 - PubMed Abstract:
Signaling molecule phosphatidylinositol 4,5-bisphosphate is produced primarily by phosphatidylinositol 4-phosphate 5-kinase (PIP5K). PIP5K is essential for the development of the human neuronal system, which has been exemplified by a recessive genetic disorder, lethal congenital contractural syndrome type 3, caused by a single aspartate-to-asparagine mutation in the kinase domain of PIP5Kγ. So far, the exact role of this aspartate residue has yet to be elucidated. In this work, we conducted structural, functional and computational studies on a zebrafish PIP5Kα variant with a mutation at the same site. Compared with the structure of the wild-type (WT) protein in the ATP-bound state, the ATP-associating glycine-rich loop of the mutant protein was severely disordered and the temperature factor of ATP was significantly higher. Both observations suggest a greater degree of disorder of the bound ATP, whereas neither the structure of the catalytic site nor the K m toward ATP was substantially affected by the mutation. Microsecond molecular dynamics simulation revealed that negative charge elimination caused by the mutation destabilized the involved hydrogen bonds and affected key electrostatic interactions in the close proximity of ATP. Taken together, our data indicated that the disease-related aspartate residue is a key node in the interaction network crucial for effective ATP binding. This work provides a paradigm of how a subtle but critical structural perturbation caused by a single mutation at the ATP-binding site abolishes the kinase activity, emphasizing that stabilizing substrate in a productive conformational state is crucial for catalysis.
- Department of Biochemistry and Molecular Biology, Michigan State University, East Lansing, MI 48824, U.S.A.
Organizational Affiliation: