An unexpected vestigial protein complex reveals the evolutionary origins of ans-triazine catabolic enzyme.
Esquirol, L., Peat, T.S., Wilding, M., Liu, J.W., French, N.G., Hartley, C.J., Onagi, H., Nebl, T., Easton, C.J., Newman, J., Scott, C.(2018) J Biological Chem 293: 7880-7891
- PubMed: 29523689
- DOI: https://doi.org/10.1074/jbc.RA118.001996
- Primary Citation Related Structures:
6C62, 6C6G - PubMed Abstract:
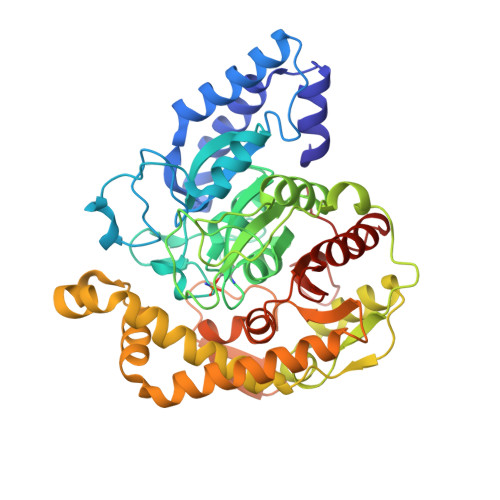
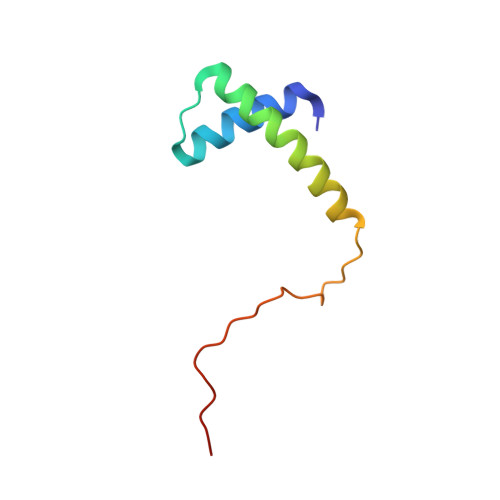
Cyanuric acid is a metabolic intermediate of s -triazines, such as atrazine (a common herbicide) and melamine (used in resins and plastics). Cyanuric acid is mineralized to ammonia and carbon dioxide by the soil bacterium Pseudomonas sp. strain ADP via three hydrolytic enzymes (AtzD, AtzE, and AtzF). Here, we report the purification and biochemical and structural characterization of AtzE. Contrary to previous reports, we found that AtzE is not a biuret amidohydrolase, but instead it catalyzes the hydrolytic deamination of 1-carboxybiuret. X-ray crystal structures of apo AtzE and AtzE bound with the suicide inhibitor phenyl phosphorodiamidate revealed that the AtzE enzyme complex consists of two independent molecules in the asymmetric unit. We also show that AtzE forms an α2β2 heterotetramer with a previously unidentified 68-amino acid-long protein (AtzG) encoded in the cyanuric acid mineralization operon from Pseudomonas sp. strain ADP. Moreover, we observed that AtzG is essential for the production of soluble, active AtzE and that this obligate interaction is a vestige of their shared evolutionary origin. We propose that AtzEG was likely recruited into the cyanuric acid-mineralizing pathway from an ancestral glutamine transamidosome that required protein-protein interactions to enforce the exclusion of solvent from the transamidation reaction.
- From the Biocatalysis and Synthetic Biology Team and.
Organizational Affiliation: