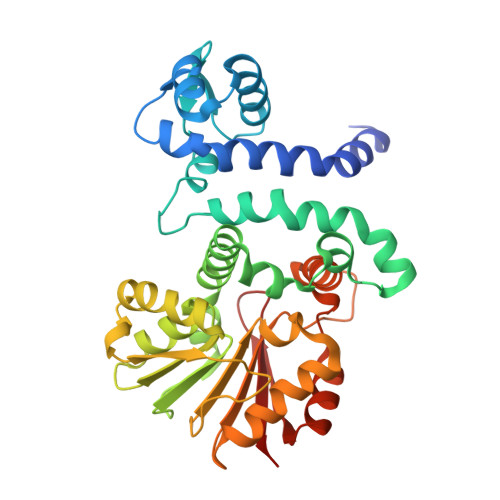
Functional and Structural Analysis of Phenazine O-Methyltransferase LaPhzM from Lysobacter antibioticus OH13 and One-Pot Enzymatic Synthesis of the Antibiotic Myxin.
Jiang, J., Guiza Beltran, D., Schacht, A., Wright, S., Zhang, L., Du, L.(2018) ACS Chem Biol 13: 1003-1012
- PubMed: 29510028 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.8b00062
- Primary Citation Related Structures:
6C5B - PubMed Abstract:
Myxin is a well-known antibiotic that had been used for decades. It belongs to the phenazine natural products that exhibit various biological activities, which are often dictated by the decorating groups on the heteroaromatic three-ring system. The three rings of myxin carry a number of decorations, including an unusual aromatic N5, N10-dioxide. We previously showed that phenazine 1,6-dicarboxylic acid (PDC) is the direct precursor of myxin, and two redox enzymes (LaPhzS and LaPhzNO1) catalyze the decarboxylative hydroxylation and aromatic N-oxidations of PDC to produce iodinin (1.6-dihydroxy- N5, N10-dioxide phenazine). In this work, we identified the LaPhzM gene from Lysobacter antibioticus OH13 and demonstrated that LaPhzM encodes a SAM-dependent O-methyltransferase converting iodinin to myxin. The results further showed that LaPhzM is responsible for both monomethoxy and dimethoxy formation in all phenazine compounds isolated from strain OH13. LaPhzM exhibits relaxed substrate selectivity, catalyzing O-methylation of phenazines with non-, mono-, or di- N-oxide. In addition, we demonstrated a one-pot biosynthesis of myxin by in vitro reconstitution of the three phenazine-ring decorating enzymes. Finally, we determined the X-ray crystal structure of LaPhzM with a bound cofactor at 1.4 Å resolution. The structure provided molecular insights into the activity and selectivity of the first characterized phenazine O-methyltransferase. These results will facilitate future exploitation of the thousands of phenazines as new antibiotics through metabolic engineering and chemoenzymatic syntheses.
- Department of Chemistry , University of Nebraska-Lincoln , Lincoln , Nebraska 68588 , United States.
Organizational Affiliation: