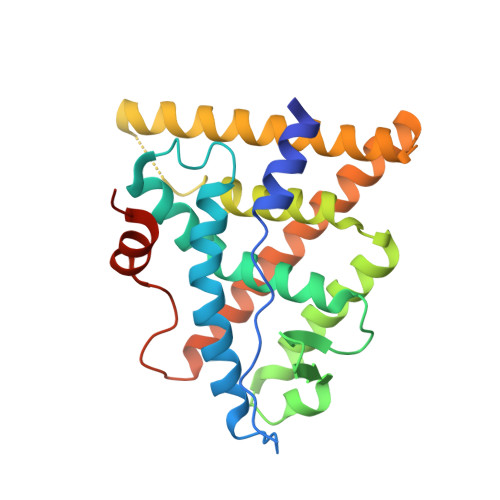
Specific stereochemistry of OP-1074 disrupts estrogen receptor alpha helix 12 and confers pure antiestrogenic activity.
Fanning, S.W., Hodges-Gallagher, L., Myles, D.C., Sun, R., Fowler, C.E., Plant, I.N., Green, B.D., Harmon, C.L., Greene, G.L., Kushner, P.J.(2018) Nat Commun 9: 2368-2368
- PubMed: 29915250 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-04413-3
- Primary Citation Related Structures:
5UFW, 5UFX, 6C42 - PubMed Abstract:
Complex tissue-specific and cell-specific signaling by the estrogen receptor (ER) frequently leads to the development of resistance to endocrine therapy for breast cancer. Pure ER antagonists, which completely lack tissue-specific agonist activity, hold promise for preventing and treating endocrine resistance, however an absence of structural information hinders the development of novel candidates. Here we synthesize a small panel of benzopyrans with variable side chains to identify pure antiestrogens in a uterotrophic assay. We identify OP-1074 as a pure antiestrogen and a selective ER degrader (PA-SERD) that is efficacious in shrinking tumors in a tamoxifen-resistant xenograft model. Biochemical and crystal structure analyses reveal a structure activity relationship implicating the importance of a stereospecific methyl on the pyrrolidine side chain of OP-1074, particularly on helix 12.
- Ben May Department for Cancer Research, University of Chicago, Chicago, IL, 60637, USA. sfanning@uchicago.edu.
Organizational Affiliation: