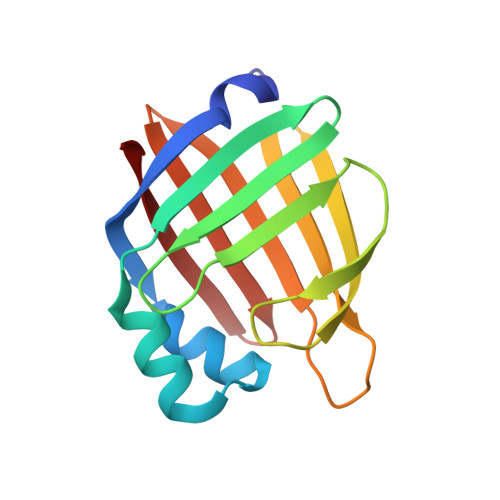
Structural characterization of life-extending Caenorhabditis elegans Lipid Binding Protein 8.
Tillman, M.C., Khadka, M., Duffy, J., Wang, M.C., Ortlund, E.A.(2019) Sci Rep 9: 9966-9966
- PubMed: 31292465
- DOI: https://doi.org/10.1038/s41598-019-46230-8
- Primary Citation Related Structures:
6C1Z - PubMed Abstract:
The lysosome plays a crucial role in the regulation of longevity. Lysosomal degradation is tightly coupled with autophagy that is induced by many longevity paradigms and required for lifespan extension. The lysosome also serves as a hub for signal transduction and regulates longevity via affecting nuclear transcription. One lysosome-to-nucleus retrograde signaling pathway is mediated by a lysosome-associated fatty acid binding protein LBP-8 in Caenorhabditis elegans. LBP-8 shuttles lysosomal lipids into the nucleus to activate lipid regulated nuclear receptors NHR-49 and NHR-80 and consequently promote longevity. However, the structural basis of LBP-8 action remains unclear. Here, we determined the first 1.3 Å high-resolution structure of this life-extending protein LBP-8, which allowed us to identify a structurally conserved nuclear localization signal and amino acids involved in lipid binding. Additionally, we described the range of fatty acids LBP-8 is capable of binding and show that it binds to life-extending ligands in worms such as oleic acid and oleoylethanolamide with high affinity.
- Department of Biochemistry, Emory University School of Medicine, 1510 Clifton Road, Atlanta, GA, 30322, USA.
Organizational Affiliation: