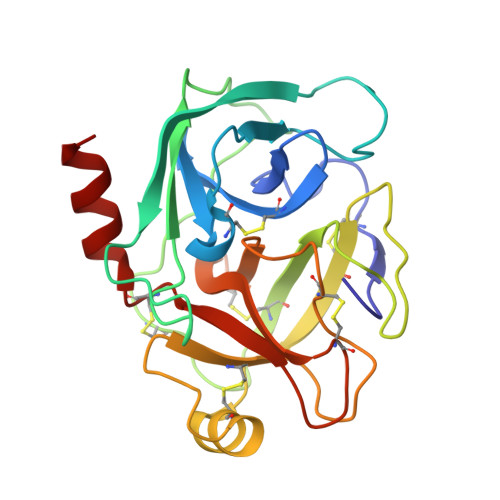
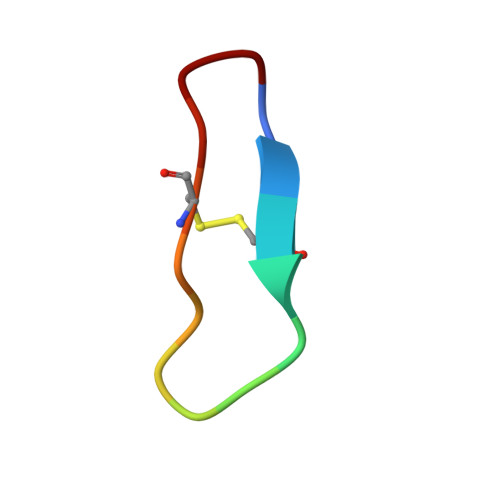
Potent, multi-target serine protease inhibition achieved by a simplified beta-sheet motif.
Chen, X., Riley, B.T., de Veer, S.J., Hoke, D.E., Van Haeften, J., Leahy, D., Swedberg, J.E., Brattsand, M., Hartfield, P.J., Buckle, A.M., Harris, J.M.(2019) PLoS One 14: e0210842-e0210842
- PubMed: 30668585 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0210842
- Primary Citation Related Structures:
6BVH - PubMed Abstract:
Engagement of an extended β-sheet is a common substrate/inhibitor interaction at the active site of serine proteases and is an important feature of Laskowski mechanism inhibitors that present a substrate-like loop to a target protease. This loop is cleaved but subsequently relegated forming a stable inhibitor/protease complex. Laskowski inhibitors are ubiquitous in nature and are used extensively in serine protease inhibitor design. However, most studies concentrate on introducing new sidechain interactions rather than the direct contributions of the substrate-like β-sheet to enzyme inhibition. Here we report the crystal structure of an simplified β-sheet inhibitory motif within the Sunflower Trypsin Inhibitor (SFTI) in complex with trypsin. We show that the intramolecular hydrogen bond network of this SFTI variant (SFTI-TCTR) engages the inhibitor sidechains that would normally interact with a target protease, giving mainchain interactions a more prominent role in complex formation. Despite having reduced sidechain interactions, this SFTI variant is remarkably potent and inhibits a diverse range of serine proteases. Crystal structural analysis and molecular modelling of SFTI-TCTR complexes again indicates an interface dominated by β-sheet interactions, highlighting the importance of this motif and the adaptability of SFTI as a scaffold for inhibitor design.
- School of Biomedical Sciences, Queensland University of Technology, Brisbane, Queensland, Australia.
Organizational Affiliation: