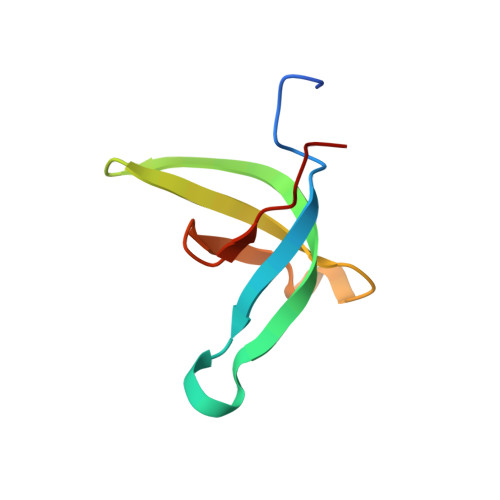
Crystal structure of chromo barrel domain of RBBP1.
Lei, M., Feng, Y., Zhou, M., Yang, Y., Loppnau, P., Li, Y., Yang, Y., Liu, Y.(2018) Biochem Biophys Res Commun 496: 1344-1348
- PubMed: 29408527 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2018.02.016
- Primary Citation Related Structures:
6BPH - PubMed Abstract:
RBBP1 is a retinoblastoma protein (pRb) binding protein acting as a repressor of gene transcription. RBBP1 is a multidomain protein including a chromo barrel domain, and its chromo barrel domain has been reported to recognize histone H4K20me3 weakly, and this binding is enhanced by the simultaneous binding of DNA. However, the molecular basis of this DNA-mediated histone binding by the chromo barrel domain of RBBP1 is unclear. Here we attempted to co-crystallize the chromo barrel domain of RBBP1 with either a histone H4K20me3 peptide alone or with both a histone H4K20me3 peptide and DNA, but only solved the peptide/DNA unbound crystal structure. Our structural analysis indicates that RBBP1 could interact with histone H4K20me3 similar to other histone binding chromo barrel domains, and the surface charge representation analysis of the chromo barrel domain of RBBP1 suggests that the chromo barrel domain of RBBP1 does not have a typical DNA binding surface, indicating that it might not bind to DNA. Consistently, our ITC assays also showed that DNA does not significantly enhance the histone binding ability of the chromo barrel domain of RBBP1.
- Hubei Key Laboratory of Genetic Regulation and Integrative Biology, School of Life Sciences, Central China Normal University, Wuhan 430079, PR China; Structural Genomics Consortium, University of Toronto, 101 College Street, Toronto, Ontario M5G 1L7, Canada.
Organizational Affiliation: