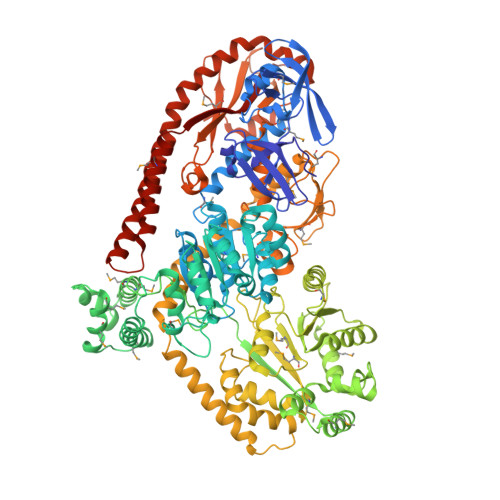
Structure of RapA, a Swi2/Snf2 protein that recycles RNA polymerase during transcription.
Shaw, G., Gan, J., Zhou, Y.N., Zhi, H., Subburaman, P., Zhang, R., Joachimiak, A., Jin, D.J., Ji, X.(2008) Structure 16: 1417-1427
- PubMed: 18786404 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2008.06.012
- Primary Citation Related Structures:
6BOG - PubMed Abstract:
RapA, as abundant as sigma70 in the cell, is an RNA polymerase (RNAP)-associated Swi2/Snf2 protein with ATPase activity. It stimulates RNAP recycling during transcription. We report a structure of RapA that is also a full-length structure for the entire Swi2/Snf2 family. RapA contains seven domains, two of which exhibit novel protein folds. Our model of RapA in complex with ATP and double-stranded DNA (dsDNA) suggests that RapA may bind to and translocate on dsDNA. Our kinetic template-switching assay shows that RapA facilitates the release of sequestered RNAP from a posttranscrption/posttermination complex for transcription reinitiation. Our in vitro competition experiment indicates that RapA binds to core RNAP only but is readily displaceable by sigma70. RapA is likely another general transcription factor, the structure of which provides a framework for future studies of this bacterial Swi2/Snf2 protein and its important roles in RNAP recycling during transcription.
- Center for Cancer Research, National Cancer Institute, National Institutes of Health, Frederick, MD 21702, USA.
Organizational Affiliation: