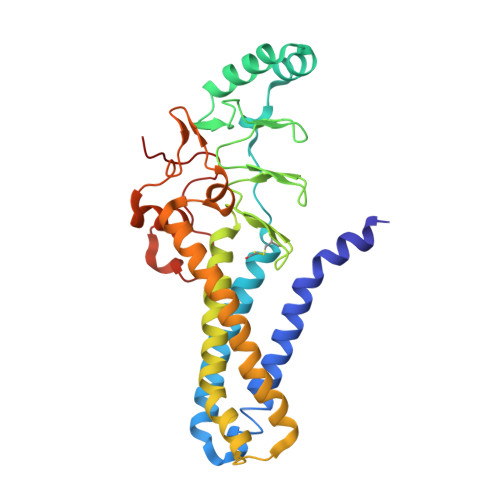
Fatty acyl recognition and transfer by an integral membraneS-acyltransferase.
Rana, M.S., Kumar, P., Lee, C.J., Verardi, R., Rajashankar, K.R., Banerjee, A.(2018) Science 359
- PubMed: 29326245 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aao6326
- Primary Citation Related Structures:
6BML, 6BMM, 6BMN, 6BMS - PubMed Abstract:
DHHC (Asp-His-His-Cys) palmitoyltransferases are eukaryotic integral membrane enzymes that catalyze protein palmitoylation, which is important in a range of physiological processes, including small guanosine triphosphatase (GTPase) signaling, cell adhesion, and neuronal receptor scaffolding. We present crystal structures of two DHHC palmitoyltransferases and a covalent intermediate mimic. The active site resides at the membrane-cytosol interface, which allows the enzyme to catalyze thioester-exchange chemistry by using fatty acyl-coenzyme A and explains why membrane-proximal cysteines are candidates for palmitoylation. The acyl chain binds in a cavity formed by the transmembrane domain. We propose a mechanism for acyl chain-length selectivity in DHHC enzymes on the basis of cavity mutants with preferences for shorter and longer acyl chains.
- Cell Biology and Neurobiology Branch, National Institutes of Child Health and Human Development, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: