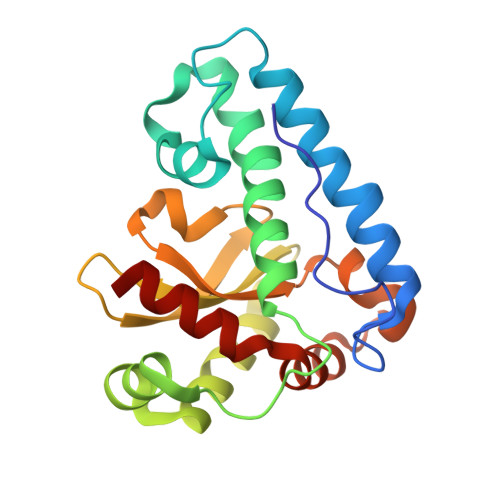
Structural characterization of a pathogenicity-related superoxide dismutase codified by a probably essential gene in Xanthomonas citri subsp. citri.
Cabrejos, D.A.L., Alexandrino, A.V., Pereira, C.M., Mendonca, D.C., Pereira, H.D., Novo-Mansur, M.T.M., Garratt, R.C., Goto, L.S.(2019) PLoS One 14: e0209988-e0209988
- PubMed: 30615696
- DOI: https://doi.org/10.1371/journal.pone.0209988
- Primary Citation Related Structures:
6BEJ - PubMed Abstract:
Citrus canker is a plant disease caused by the bacteria Xanthomonas citri subsp. citri that affects all domestic varieties of citrus. Some annotated genes from the X. citri subsp. citri genome are assigned to an interesting class named "pathogenicity, virulence and adaptation". Amongst these is sodM, which encodes for the gene product XcSOD, one of four superoxide dismutase homologs predicted from the genome. SODs are widespread enzymes that play roles in the oxidative stress response, catalyzing the degradation of the deleterious superoxide radical. In Xanthomonas, SOD has been associated with pathogenesis as a counter measure against the plant defense response. In this work we initially present the 1.8 Å crystal structure of XcSOD, a manganese containing superoxide dismutase from Xanthomonas citri subsp. citri. The structure bears all the hallmarks of a dimeric member of the MnSOD family, including the conserved hydrogen-bonding network residues. Despite the apparent gene redundancy, several attempts to obtain a sodM deletion mutant were unsuccessful, suggesting the encoded protein to be essential for bacterial survival. This intriguing observation led us to extend our structural studies to the remaining three SOD homologs, for which comparative models were built. The models imply that X. citri subsp. citri produces an iron-containing SOD which is unlikely to be catalytically active along with two conventional Cu,ZnSODs. Although the latter are expected to possess catalytic activity, we propose they may not be able to replace XcSOD for reasons such as distinct subcellular compartmentalization or differential gene expression in pathogenicity-inducing conditions.
- Laboratório de Biologia Estrutural, Grupo de Cristalografia, Instituto de Física de São Carlos, Universidade de São Paulo, São Carlos, SP, Brazil.
Organizational Affiliation: