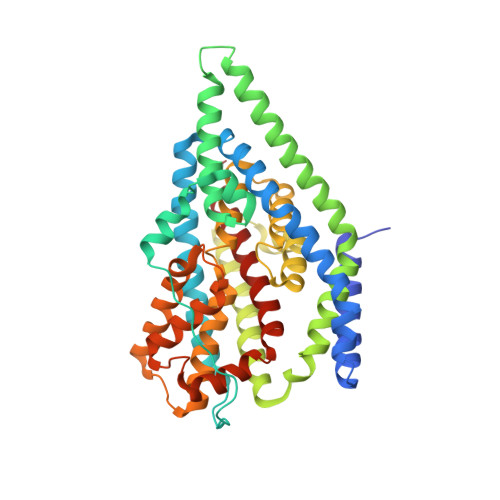
Structural characterisation reveals insights into substrate recognition by the glutamine transporter ASCT2/SLC1A5.
Scopelliti, A.J., Font, J., Vandenberg, R.J., Boudker, O., Ryan, R.M.(2018) Nat Commun 9: 38-38
- PubMed: 29295993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02444-w
- Primary Citation Related Structures:
6BAT, 6BAU, 6BAV, 6BMI - PubMed Abstract:
Cancer cells undergo a shift in metabolism where they become reliant on nutrients such as the amino-acid glutamine. Glutamine enters the cell via the alanine/serine/cysteine transporter 2 (ASCT2) that is upregulated in several cancers to maintain an increased supply of this nutrient and are therefore an attractive target in cancer therapeutic development. ASCT2 belongs to the glutamate transporter (SLC1A) family but is the only transporter in this family able to transport glutamine. The structural basis for glutamine selectivity of ASCT2 is unknown. Here, we identify two amino-acid residues in the substrate-binding site that are responsible for conferring glutamine selectivity. We introduce corresponding mutations into a prokaryotic homologue of ASCT2 and solve four crystal structures, which reveal the structural basis for neutral amino acid and inhibitor binding in this family. This structural model of ASCT2 may provide a basis for future development of selective ASCT2 inhibitors to treat glutamine-dependent cancers.
- Transporter Biology Group, Discipline of Pharmacology, Sydney Medical School, University of Sydney, Sydney, NSW, 2006, Australia.
Organizational Affiliation: