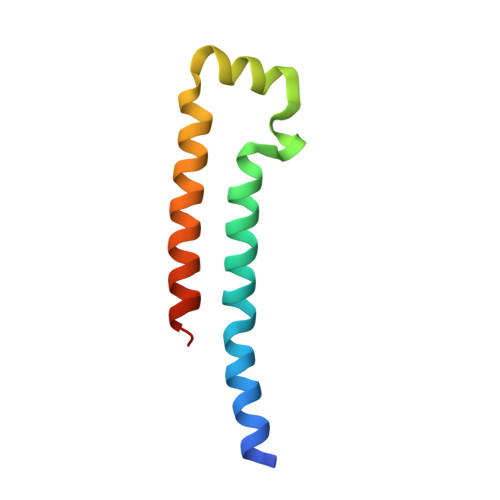
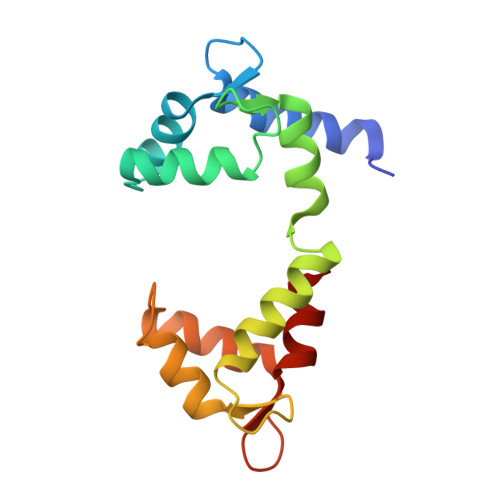
A Calmodulin C-Lobe Ca
Chang, A., Abderemane-Ali, F., Hura, G.L., Rossen, N.D., Gate, R.E., Minor, D.L.(2018) Neuron 97: 836-852.e6
- PubMed: 29429937 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.neuron.2018.01.035
- Primary Citation Related Structures:
6B8L, 6B8M, 6B8N, 6B8P, 6B8Q - PubMed Abstract:
Kv7 (KCNQ) voltage-gated potassium channels control excitability in the brain, heart, and ear. Calmodulin (CaM) is crucial for Kv7 function, but how this calcium sensor affects activity has remained unclear. Here, we present X-ray crystallographic analysis of CaM:Kv7.4 and CaM:Kv7.5 AB domain complexes that reveal an Apo/CaM clamp conformation and calcium binding preferences. These structures, combined with small-angle X-ray scattering, biochemical, and functional studies, establish a regulatory mechanism for Kv7 CaM modulation based on a common architecture in which a CaM C-lobe calcium-dependent switch releases a shared Apo/CaM clamp conformation. This C-lobe switch inhibits voltage-dependent activation of Kv7.4 and Kv7.5 but facilitates Kv7.1, demonstrating that mechanism is shared by Kv7 isoforms despite the different directions of CaM modulation. Our findings provide a unified framework for understanding how CaM controls different Kv7 isoforms and highlight the role of membrane proximal domains for controlling voltage-gated channel function. VIDEO ABSTRACT.
- Cardiovascular Research Institute, University of California San Francisco, San Francisco, CA 94158, USA.
Organizational Affiliation: