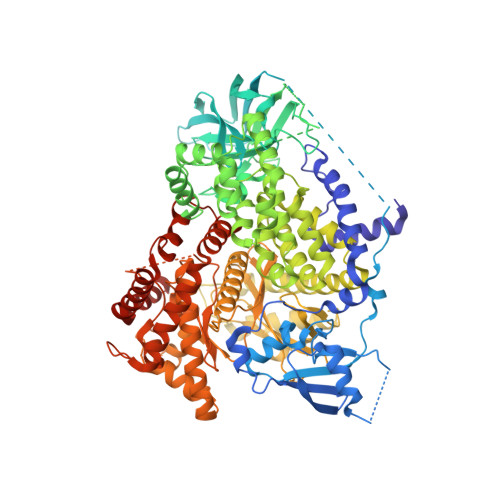
Design of Selective Benzoxazepin PI3K delta Inhibitors Through Control of Dihedral Angles.
Safina, B.S., Elliott, R.L., Forrest, A.K., Heald, R.A., Murray, J.M., Nonomiya, J., Pang, J., Salphati, L., Seward, E.M., Staben, S.T., Ultsch, M., Wei, B., Yang, W., Sutherlin, D.P.(2017) ACS Med Chem Lett 8: 936-940
- PubMed: 28947940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.7b00170
- Primary Citation Related Structures:
6AUD - PubMed Abstract:
A novel selective benzoxazepin inhibitor of PI3Kδ has been discovered. Beginning from compound 3 , an αPI3K inhibitor, we utilized structure-based drug design and computational analysis of dihedral torsion angles to optimize for PI3Kδ isoform potency and isoform selectivity. Further medicinal chemistry optimization of the series led to the identification of 24 , a highly potent and selective inhibitor of PI3Kδ.
- Discovery Chemistry, Genentech, Inc., 1 DNA Way, South San Francisco, California 94080, United States.
Organizational Affiliation: