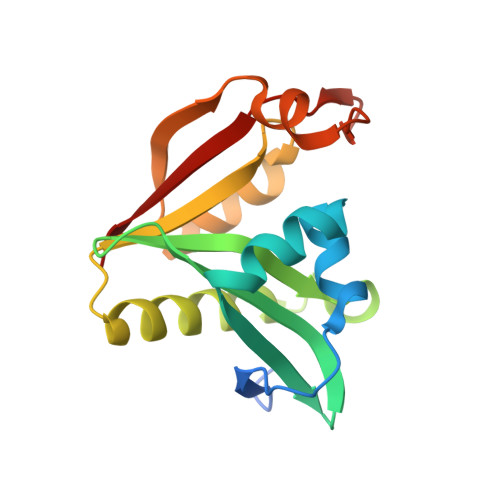
Structural characterization of a GNAT family acetyltransferase from Elizabethkingia anophelis bound to acetyl-CoA reveals a new dimeric interface.
Shirmast, P., Ghafoori, S.M., Irwin, R.M., Abendroth, J., Mayclin, S.J., Lorimer, D.D., Edwards, T.E., Forwood, J.K.(2021) Sci Rep 11: 1274-1274
- PubMed: 33446675
- DOI: https://doi.org/10.1038/s41598-020-79649-5
- Primary Citation Related Structures:
6AO7 - PubMed Abstract:
General control non-repressible 5 (GCN5)-related N-acetyltransferases (GNATs) catalyse the acetylation of a diverse range of substrates, thereby orchestrating a variety of biological processes within prokaryotes and eukaryotes. GNAT enzymes can catalyze the transfer of an acetyl group from acetyl coenzyme A to substrates such as aminoglycoside antibiotics, amino acids, polyamines, peptides, vitamins, catecholamines, and large macromolecules including proteins. Although GNATs generally exhibit low to moderate sequence identity, they share a conserved catalytic fold and conserved structural motifs. In this current study we characterize the high-resolution X-ray crystallographic structure of a GNAT enzyme bound with acetyl-CoA from Elizabethkingia anophelis, an important multi-drug resistant bacterium. The tertiary structure is comprised of six α-helices and nine β-strands, and is similar with other GNATs. We identify a new and uncharacterized GNAT dimer interface, which is conserved in at least two other unpublished GNAT structures. This suggests that GNAT enzymes can form at least five different types of dimers, in addition to a range of other oligomers including trimer, tetramer, hexamer, and dodecamer assemblies. The high-resolution structure presented in this study is suitable for future in-silico docking and structure-activity relationship studies.
- School of Biomedical Sciences, Charles Sturt University, Wagga Wagga, NSW, 2678, Australia.
Organizational Affiliation: