Structure-Activity Relationship Study of Novel 6-Aryl-2-benzoyl-pyridines as Tubulin Polymerization Inhibitors with Potent Antiproliferative Properties.
Chen, H., Deng, S., Wang, Y., Albadari, N., Kumar, G., Ma, D., Li, W., White, S.W., Miller, D.D., Li, W.(2020) J Med Chem 63: 827-846
- PubMed: 31860298 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01815
- Primary Citation Related Structures:
6AGK, 6PC4 - PubMed Abstract:
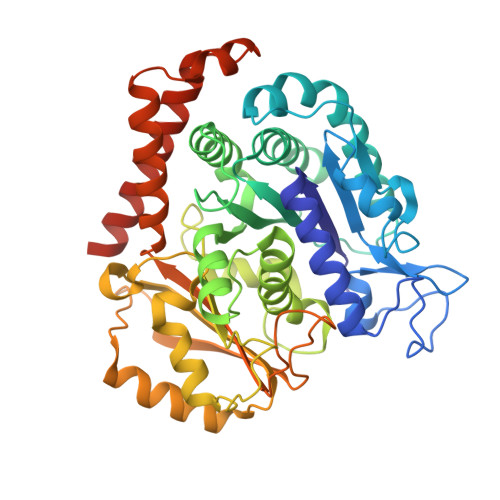
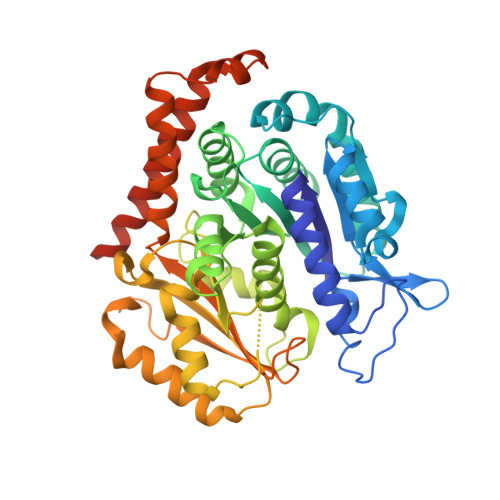
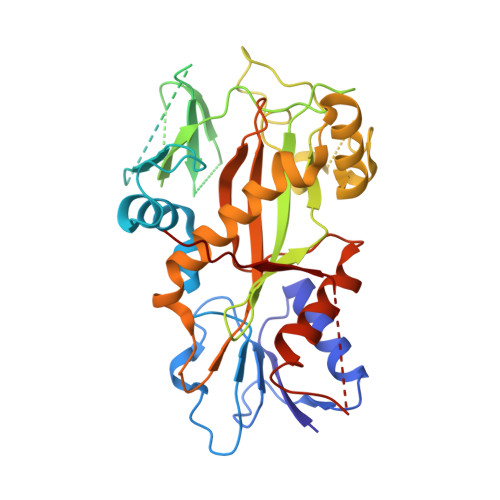
We recently reported the crystal structure of tubulin in complex with a colchicine binding site inhibitor (CBSI), ABI-231, having 2-aryl-4-benzoyl-imidazole (ABI). Based on this and additional crystal structures, here we report the structure-activity relationship study of a novel series of pyridine analogues of ABI-231, with compound 4v being the most potent one (average IC 50 ∼ 1.8 nM) against a panel of cancer cell lines. We determined the crystal structures of another potent CBSI ABI-274 and 4v in complex with tubulin and confirmed their direct binding at the colchicine site. 4v inhibited tubulin polymerization, strongly suppressed A375 melanoma tumor growth, induced tumor necrosis, disrupted tumor angiogenesis, and led to tumor cell apoptosis in vivo. Collectively, these studies suggest that 4v represents a promising new generation of tubulin inhibitors.
- Department of Pharmaceutical Sciences, College of Pharmacy , University of Tennessee Health Science Center , Memphis , Tennessee 38163 , United States.
Organizational Affiliation: