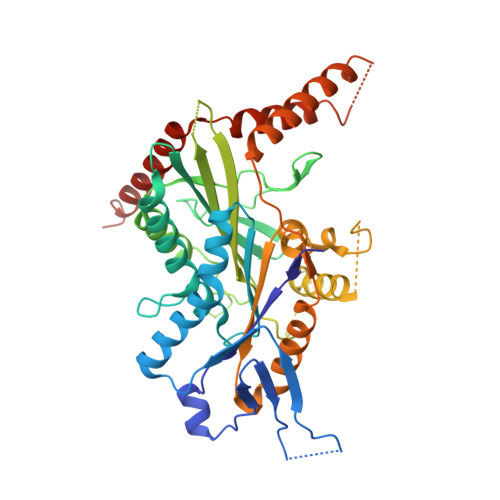
Coiled-coil 1-mediated fastening of the neck and motor domains for kinesin-3 autoinhibition.
Ren, J.Q., Wang, S., Chen, H., Wang, W.J., Huo, L., Feng, W.(2018) Proc Natl Acad Sci U S A 115: E11933-E11942
- PubMed: 30463954 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1811209115
- Primary Citation Related Structures:
6A1Z, 6A20 - PubMed Abstract:
In kinesin-3, the coiled-coil 1 (CC1) can sequester the preceding neck coil (NC) for autoinhibition, but the underlying mechanism is poorly understood. Here, we determined the structures of the uninhibited motor domain (MD)-NC dimer and inhibited MD-NC-CC1 monomer of kinesin-3 KIF13B. In the MD-NC-CC1 monomer, CC1 is broken into two short helices that unexpectedly interact with both the NC and the MD. Compared with the MD-NC dimer, the CC1-mediated integration of NC and MD not only blocks the NC dimer formation, but also prevents the neck linker (NL) undocking and the ADP release from the MD. Mutations of the essential residues in the interdomain interaction interface in the MD-NC-CC1 monomer restored the MD activity. Thus, CC1 fastens the neck domain and MD and inhibits both NC and NL. This CC1-mediated lockdown of the entire neck domain may represent a paradigm for kinesin autoinhibition that could be applicable to other kinesin-3 motors.
- National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 100101 Beijing, China.
Organizational Affiliation: