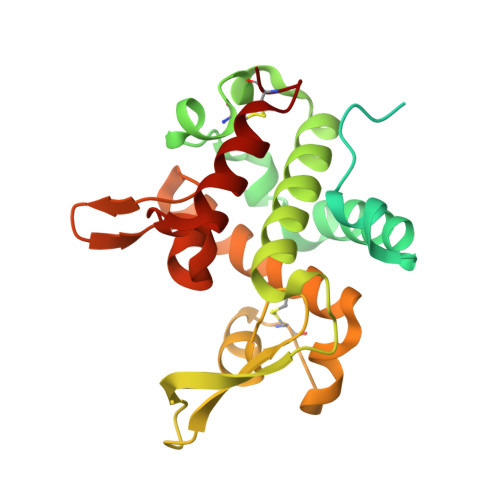
A lysozyme with altered substrate specificity facilitates prey cell exit by the periplasmic predator Bdellovibrio bacteriovorus.
Harding, C.J., Huwiler, S.G., Somers, H., Lambert, C., Ray, L.J., Till, R., Taylor, G., Moynihan, P.J., Sockett, R.E., Lovering, A.L.(2020) Nat Commun 11: 4817-4817
- PubMed: 32968056 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-18139-8
- Primary Citation Related Structures:
6TA9, 6TAB, 6TAD, 6TAF - PubMed Abstract:
Lysozymes are among the best-characterized enzymes, acting upon the cell wall substrate peptidoglycan. Here, examining the invasive bacterial periplasmic predator Bdellovibrio bacteriovorus, we report a diversified lysozyme, DslA, which acts, unusually, upon (GlcNAc-) deacetylated peptidoglycan. B. bacteriovorus are known to deacetylate the peptidoglycan of the prey bacterium, generating an important chemical difference between prey and self walls and implying usage of a putative deacetyl-specific "exit enzyme". DslA performs this role, and ΔDslA strains exhibit a delay in leaving from prey. The structure of DslA reveals a modified lysozyme superfamily fold, with several adaptations. Biochemical assays confirm DslA specificity for deacetylated cell wall, and usage of two glutamate residues for catalysis. Exogenous DslA, added ex vivo, is able to prematurely liberate B. bacteriovorus from prey, part-way through the predatory lifecycle. We define a mechanism for specificity that invokes steric selection, and use the resultant motif to identify wider DslA homologues.
- Institute for Microbiology and Infection, School of Biosciences, University of Birmingham, Birmingham, B15 2TT, UK.
Organizational Affiliation: