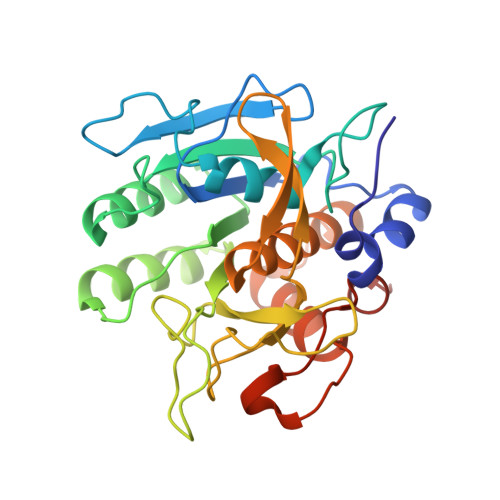
Insight into subtilisin E-S7 cleavage pattern based on crystal structure and hydrolysates peptide analysis.
Tang, H., Zhang, J., Shi, K., Aihara, H., Du, G.(2019) Biochem Biophys Res Commun 512: 623-628
- PubMed: 30914195 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrc.2019.03.064
- Primary Citation Related Structures:
6O44 - PubMed Abstract:
The X-ray crystallographic structure of the mature form of subtilisin E-S7 (SES7) at 1.90 Å resolution is reported here. Structural comparisons between the previously reported propeptide-subtilisin E complex (1SCJ) and our mature form subtilisin E-S7 (6O44) provide insight into active site adjustments involved in catalysis and specificity. To further investigate the protease substrate selectivity mechanism, we used SES7 to hydrolyze skim milk and analyzed the hydrolysates by LC-MS for peptide identification. The cleavage pattern suggests a high preference for proline at substrate P2 position. The results based on the peptide analysis are consistent with our structural observations, which is instrumental in future protein engineering by rational design. Furthermore, the ACE-inhibitor and NLN-inhibitor activity of the hydrolysates were determined to assess the utility of SES7 for further industrial applications; IC 50 -ACE = 67 ± 0.92 μg/mL and IC 50 -NLN = 263 ± 13 μg/mL.
- Key Laboratory of Industrial Biotechnology, Ministry of Education, School of Biotechnology, Jiangnan University, 1800 Lihu Road, Wuxi, 214122, Jiangsu, China; School of Biotechnology, Jiangnan University, 1800 Lihu Road, Wuxi, 214122, Jiangsu, China.
Organizational Affiliation: