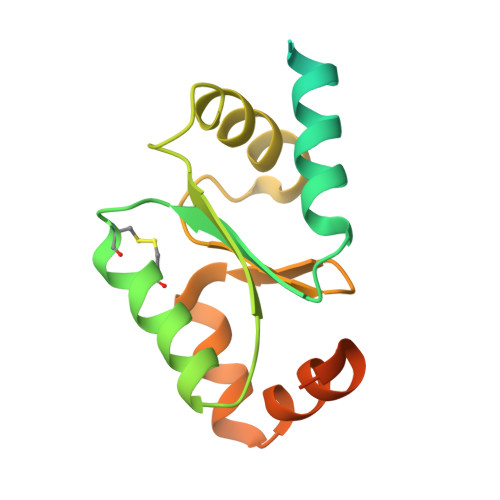
Crystal Structure of Wheat Glutaredoxin and Its Application in Improving the Processing Quality of Flour.
Sun, X., Chen, M., Jia, F., Hou, Y., Hu, S.Q.(2018) J Agric Food Chem 66: 12079-12087
- PubMed: 30346751 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jafc.8b03590
- Primary Citation Related Structures:
5ZVL - PubMed Abstract:
Glutaredoxin (Grx) is a ubiquitous oxidoreductase that plays a vital role in maintaining cellular redox homeostasis. In comparison to Grx from other organisms, plant Grx is unique in that it has many isoforms, which, thus, suggests probably diverse functions and mechanisms. Therefore, structure-function characterization of plant Grx is necessary to have in-depth knowledge and explore its application in industry. In this study, wheat Grx (wGrx) was overexpressed and purified and the crystal structure of wGrx was determined at 2.94 Å resolution. Interestingly, the structure for the first time captured both the oxidized form and the transient state of reduced-oxidized wGrx in a crystal. The mutagenesis of wGrx suggests that it adopts a monothiol catalytic mechanism. wGrx has the ability to reduce wheat thioredoxin (wTrx), and this is the first example of the reduction of thioredoxin subgroup h class II by Grx. Flour farinograph and dynamic rheological analysis showed that wGrx together with wTrx has a positive effect on dough formation, which is probably attributed to the increased sodium dodecyl sulfate (SDS)-insoluble gluten macropolymer (GMP) through increasing the intermolecular disulfide bond induced by the wGrx-wTrx system. The results indicate great potential of wGrx-wTrx as a novel synergetic enzymatic additive and may be employed to fine-tune the processing performance of food related to the redox reaction.
- School of Food Science and Engineering , South China University of Technology , Guangzhou , Guangdong 510641 , People's Republic of China.
Organizational Affiliation: