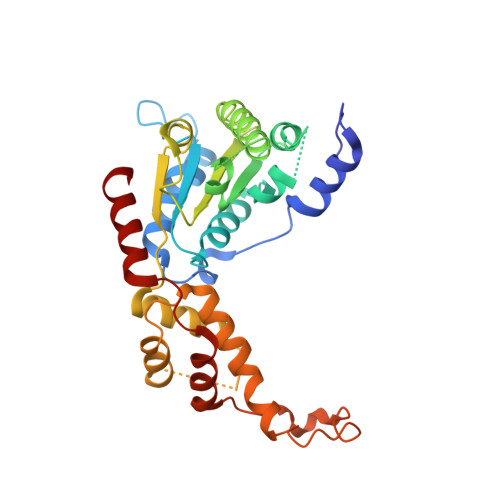
Structural and Molecular Basis for Katanin-Mediated Severing of Glutamylated Microtubules.
Shin, S.C., Im, S.K., Jang, E.H., Jin, K.S., Hur, E.M., Kim, E.E.(2019) Cell Rep 26: 1357-1367.e5
- PubMed: 30699360 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2019.01.020
- Primary Citation Related Structures:
5ZQL, 5ZQM - PubMed Abstract:
Katanin was the first microtubule (MT)-severing enzyme discovered, but how katanin executes MT severing remains poorly understood. Here, we report X-ray crystal structures of the apo and ATPγS-bound states of the catalytic AAA domain of human katanin p60 at 3.0 and 2.9 Å resolution, respectively. Comparison of the two structures reveals conformational changes induced by ATP binding and how such changes ensure hexamer stability. Moreover, we uncover structural details of pore loops (PLs) and show that Arg283, a residue unique to katanin among MT-severing enzymes, protrudes from PL1 and lines the entry of the catalytic pore. Functional studies suggest that PL1 and Arg283 play essential roles in the recognition and remodeling of the glutamylated, C-terminal tubulin tail and regulation of axon growth. In addition, domain-swapping experiments in katanin and spastin suggest that the non-homologous N-terminal region, which contains the MT-interacting and trafficking domain and a linker, confers specificity to the severing process.
- Biomedical Research Institute, Korea Institute of Science and Technology, Seoul, 02792, Republic of Korea.
Organizational Affiliation: