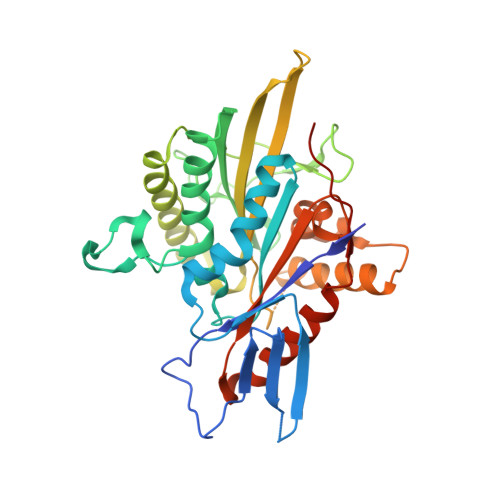
Structural and Thermodynamic Basis of the Enhanced Interaction between Kinesin Spindle Protein Eg5 and STLC-type Inhibitors.
Yokoyama, H., Sawada, J.I., Sato, K., Ogo, N., Kamei, N., Ishikawa, Y., Hara, K., Asai, A., Hashimoto, H.(2018) ACS Omega 3: 12284-12294
- PubMed: 31459302 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsomega.8b00778
- Primary Citation Related Structures:
5ZO7, 5ZO8, 5ZO9 - PubMed Abstract:
For a better understanding of protein-inhibitor interactions, we report structural, thermodynamic, and biological analyses of the interactions between S -trityl-l-cysteine (STLC) derivatives and the motor domain of kinesin spindle protein Eg5. Binding of STLC-type inhibitors to Eg5 was enthalpically driven and entropically unfavorable. The introduction of a para -methoxy substituent in one phenyl ring of STLC enhances its inhibitory activity resulting from a larger enthalpy gain possibly due to the increased shape complementarity. The substituent fits to a recess in the binding pocket. To avoid steric hindrance, the substituted STLC is nudged toward the side opposite to the recess, which enhances the interaction of Eg5 with the remaining part of the inhibitor. Further introduction of an ethylene linkage between two phenyl rings enhances Eg5 inhibitory activity by reducing the loss of entropy in forming the complex. This study provides valuable examples of enhancing protein-inhibitor interactions without forming additional hydrogen bonds.
- Faculty of Pharmaceutical Sciences, Tokyo University of Science, 2641 Yamazaki, Noda, Chiba 278-8510, Japan.
Organizational Affiliation: