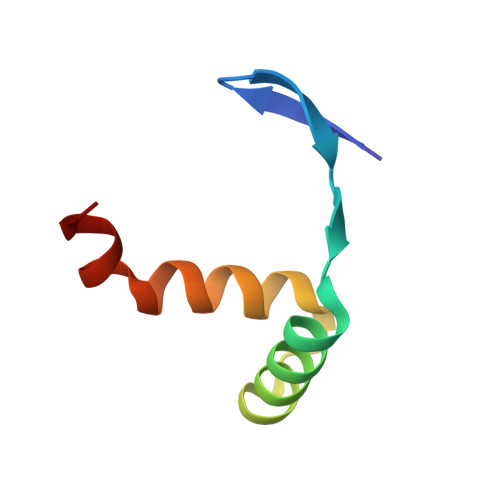
The crystal structure of the TCP domain of PCF6 in Oryza sativa L. reveals an RHH-like fold.
Sun, L., Zou, X., Jiang, M., Wu, X., Chen, Y., Wang, Q., Wang, Q., Chen, L., Wu, Y.(2020) FEBS Lett 594: 1296-1306
- PubMed: 31898812 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13727
- Primary Citation Related Structures:
5ZKT - PubMed Abstract:
The Teosinte branched 1/Cycloidea/Proliferating cell factor (TCP) domain is an evolutionarily conserved DNA binding domain unique to the plant kingdom. To date, the functions of TCPs have been well studied, but the three-dimensional structure of the TCP domain is lacking. Here, we have determined the crystal structure of the TCP domain from OsPCF6. The structure reveals that the TCP domain adopts three short β-strands followed by a helix-loop-helix structure, distinct from the canonical basic helix-loop-helix structure. This folded domain shows high structural similarity to the ribbon-helix-helix (RHH) transcriptional repressors, a family of DNA binding proteins with a conserved 3D structural motif (RHH fold), indicating that TCPs could be reclassified as RHH proteins. Our work will provide insight toward a better understanding of the mechanisms underlying TCP protein function.
- Provincial University Key Laboratory of Cellular Stress Response and Metabolic Regulation, College of Life Science, Fujian Normal University, Fuzhou, China.
Organizational Affiliation: