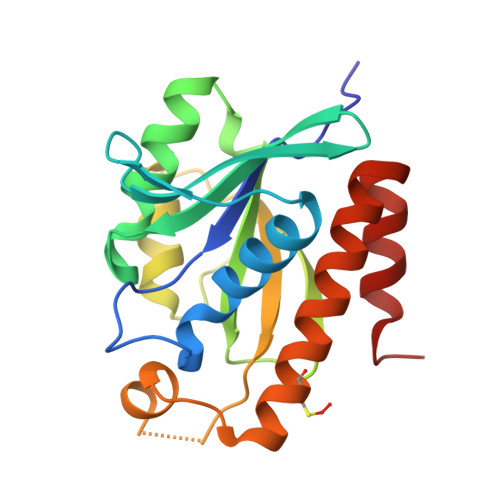
Role of methionine 71 in substrate recognition and structural integrity of bacterial peptidyl-tRNA hydrolase.
Shahid, S., Kabra, A., Mundra, S., Pal, R.K., Tripathi, S., Jain, A., Arora, A.(2018) Biochim Biophys Acta 1866: 865-874
- PubMed: 29733913
- DOI: https://doi.org/10.1016/j.bbapap.2018.05.002
- Primary Citation Related Structures:
5ZK0 - PubMed Abstract:
Bacterial peptidyl-tRNA hydrolase (Pth) is an essential enzyme that alleviates tRNA starvation by recycling prematurely dissociated peptidyl-tRNAs. The specificity of Pth for N-blocked-aminoacyl-tRNA has been proposed to be contingent upon conserved residue N14 forming a hydrogen bond with the carbonyl of the first peptide bond in the substrate. M71 is involved in forming a conserved hydrogen bond with N14. Other interactions facilitating this recognition are not known. The structure, dynamics, and stability of the M71A mutant of Pth from Vibrio cholerae (VcPth) were characterized by X-ray crystallography, NMR spectroscopy, MD simulations and DSC. Crystal structure of M71A mutant was determined. In the structure, the dimer interface is formed by the insertion of six C-terminal residues of one molecule into the active site of another molecule. The side-chain amide of N14 was hydrogen bonded to the carbonyl of the last peptide bond formed between residues A196 and E197, and also to A71. The CSP profile of mutation was similar to that observed for the N14D mutant. M71A mutation lowered the thermal stability of the protein. Our results indicate that the interactions of M71 with N14 and H24 play an important role in optimal positioning of their side-chains relative to the peptidyl-tRNA substrate. Overall, these interactions of M71 are important for the activity, stability, and compactness of the protein. The work presented provides original and new structural and dynamics information that significantly enhances our understanding of the network of interactions that govern this enzyme's activity and selectivity.
- Molecular and Structural Biology Division, CSIR-Central Drug Research Institute, Lucknow 226031, India.
Organizational Affiliation: