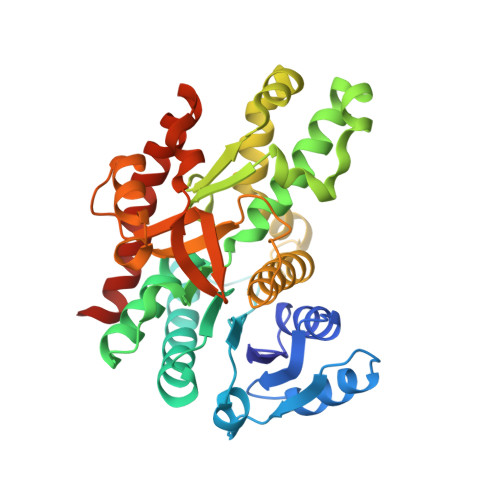
Structure of glyoxysomal malate dehydrogenase (MDH3) from Saccharomyces cerevisiae.
Moriyama, S., Nishio, K., Mizushima, T.(2018) Acta Crystallogr F Struct Biol Commun 74: 617-624
- PubMed: 30279312 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18011895
- Primary Citation Related Structures:
5ZI2, 5ZI3, 5ZI4 - PubMed Abstract:
Malate dehydrogenase (MDH), a carbohydrate and energy metabolism enzyme in eukaryotes, catalyzes the interconversion of malate to oxaloacetate (OAA) in conjunction with that of nicotinamide adenine dinucleotide (NAD + ) to NADH. Three isozymes of MDH have been reported in Saccharomyces cerevisiae: MDH1, MDH2 and MDH3. MDH1 is a mitochondrial enzyme and a member of the tricarboxylic acid cycle, whereas MDH2 is a cytosolic enzyme that functions in the glyoxylate cycle. MDH3 is a glyoxysomal enzyme that is involved in the reoxidation of NADH, which is produced during fatty-acid β-oxidation. The affinity of MDH3 for OAA is lower than those of MDH1 and MDH2. Here, the crystal structures of yeast apo MDH3, the MDH3-NAD + complex and the MDH3-NAD + -OAA ternary complex were determined. The structure of the ternary complex suggests that the active-site loop is in the open conformation, differing from the closed conformations in mitochondrial and cytosolic malate dehydrogenases.
- Picobiology Institute, Graduate School of Life Science, University of Hyogo, 3-2-1 Kouto, Kamigori-cho, Ako-gun, Hyogo 678-1297, Japan.
Organizational Affiliation: