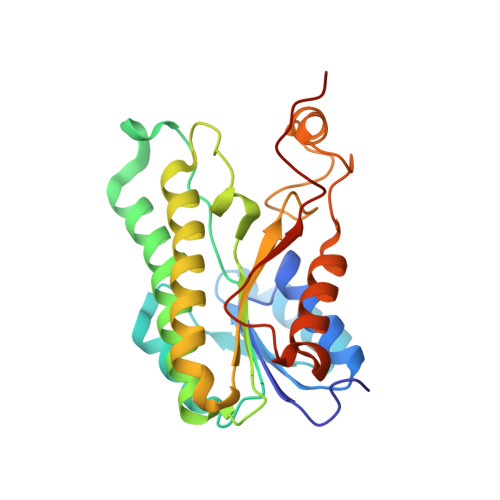
A potential substrate binding pocket of BdcA plays a critical role in NADPH recognition and biofilm dispersal
Yang, W.S., Hong, Y., Zhang, Y., Wang, D.C., Li, D.F., Hou, Y.J.(2018) Biochem Biophys Res Commun 497: 863-868
- PubMed: 29462616 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2018.02.143
- Primary Citation Related Structures:
5Z2L - PubMed Abstract:
Biofilm dispersal is characterized by the cell detachment from biofilms and expected to provide novel "anti-biofilm" approaches of prevention and treatment of biofilms in clinical and industrial settings. The E.coli protein BdcA has been identified as a biofilm dispersal factor and designed to be an important component in engineered applications to control biofilm formation. It belongs to short-chain dehydrogenase/reductase (SDR) family with the specific affinity to NADPH. Here, we show the structure of BdcA in complex with NADPH and confirm that NADPH binding is requisite for BdcA facilitating cell motility and increasing biofilm dispersal. Especially, we observe a potential substrate binding pocket surrounded by hydrophobic residues upon NADPH binding and present evidences that this pocket is essential for BdcA binding NADPH and exerting its biological functions. Our study provides the clues for illuminating the molecular mechanism of BdcA regulating biofilm dispersal and better utilizing BdcA to eliminate the biofilms.
- National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China; University of Chinese Academy of Sciences, Beijing 100049, China.
Organizational Affiliation: