A proposed role for MSC to reserve the canonical function in high eukaryotes prior to stimuli
Zheng, L., Ali, H., Wang, J., Guo, M., Fang, P.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Lysine--tRNA ligase, heat inducible | 525 | Escherichia coli K-12 | Mutation(s): 1 Gene Names: lysU, b4129, JW4090 EC: 6.1.1.6 | 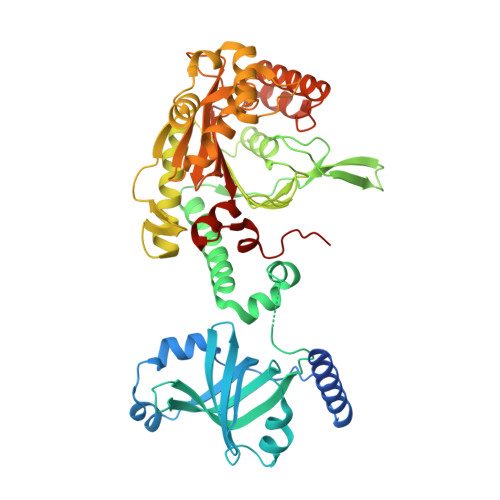 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A8N5 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| B4P Download:Ideal Coordinates CCD File | D [auth A], H [auth B], L [auth C] | BIS(ADENOSINE)-5'-TETRAPHOSPHATE C20 H28 N10 O19 P4 YOAHKNVSNCMZGQ-XPWFQUROSA-N |  | ||
| CA Download:Ideal Coordinates CCD File | E [auth A] F [auth A] G [auth A] I [auth B] J [auth B] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 144.544 | α = 90 |
| b = 249.855 | β = 90 |
| c = 179.652 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-3000 | data reduction |
| MOLREP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health | United States | GM100136 |
| National Institutes of Health | United States | GM106134 |
| Shanghai Pujiang Program | China | 17PJ1410800 |
| National Natural Science Foundation of China | China | 21778067 |
| National Natural Science Foundation of China | China | 21778064 |