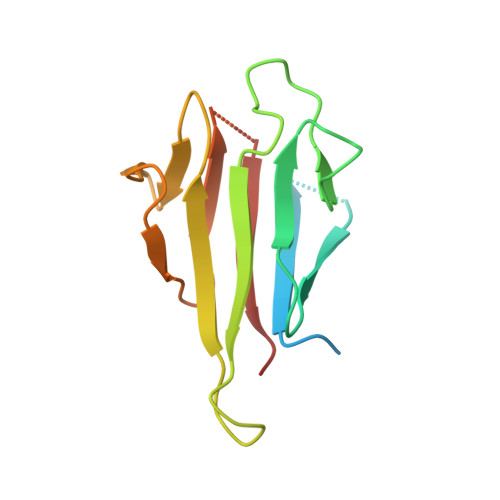
Structural insights into the recognition of phosphorylated Hop1 by Mek1
Xie, C., He, C., Jiang, Y., Yu, H., Cheng, L., Nshogoza, G., Ala, M.S., Tian, C., Wu, J., Shi, Y., Li, F.(2018) Acta Crystallogr D Struct Biol 74: 1027-1038
- PubMed: 30289413
- DOI: https://doi.org/10.1107/S2059798318011993
- Primary Citation Related Structures:
5YYX, 5YYZ - PubMed Abstract:
The FHA domain-containing protein Mek1 is a meiosis-specific kinase that is involved in the regulation of interhomolog recombination in meiosis in Saccharomyces cerevisiae. The recruitment and activation of Mek1 require the phosphorylation of the chromosome axis protein Hop1 at Thr318 (pT318), which is necessary for recognition by the Mek1 FHA domain. Here, crystal structures of the Mek1 FHA domain in the apo state and in complex with the Hop1 pT318 peptide are presented, demonstrating that the hydrophobic residues Phe320 and Val321 at the pT+2 and pT+3 positions in the ligand contribute to the preferential recognition. It was further found that in Schizosaccharomyces pombe Mek1 FHA binds both pT15 in its N-terminal SQ/TQ cluster domain (SCD) and pT270 in the Hop1 SCD. The results revealed the structural basis for the preferential recognition of phosphorylated Hop1 by Mek1 in S. cerevisiae and facilitate the understanding of the interaction between the S. pombe Mek1 FHA domain and its binding targets.
- High Magnetic Field Laboratory, Chinese Academy of Sciences, 50 Shushanhu Road, Hefei, Anhui 230031, People's Republic of China.
Organizational Affiliation: