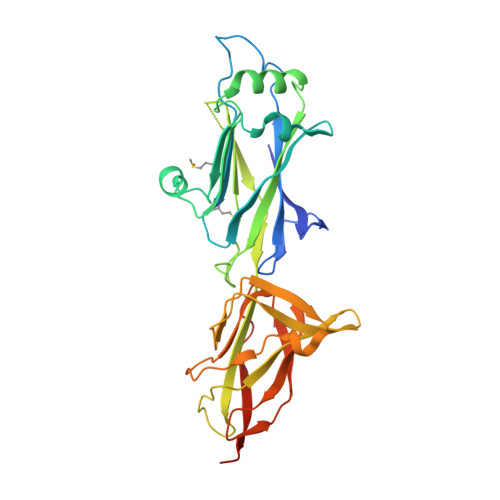
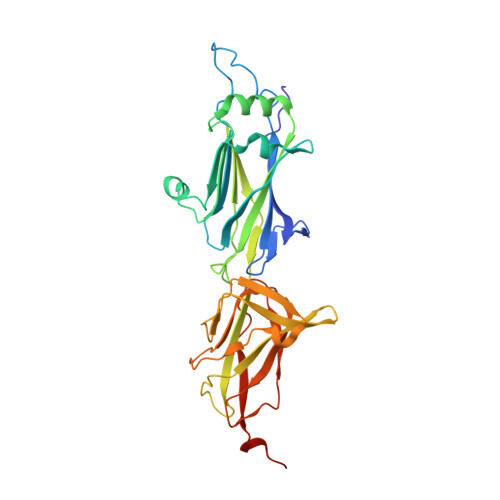
Bent conformation of a backbone pilin N-terminal domain supports a three-stage pilus assembly mechanism.
Chaurasia, P., Pratap, S., Palva, A., von Ossowski, I., Krishnan, V.(2018) Commun Biol 1: 94-94
- PubMed: 30271975 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-018-0100-0
- Primary Citation Related Structures:
5YU5, 5YXG, 5YXO, 5Z0Z, 5Z24 - PubMed Abstract:
Effective colonization of host cells by some Gram-positive bacteria often involves using lengthy, adhesive macromolecular structures called sortase-dependent pili. Among commensals, the gut-adapted Lactobacillus rhamnosus GG strain encodes the operons for two varieties of these pili (SpaCBA and SpaFED), with each structure consisting of backbone, tip, and basal pilin subunits. Although the tertiary structure was recently solved for the backbone subunit (SpaA) of the SpaCBA pilus, no structural information exists for its counterpart in the SpaFED pilus. Here, we report several crystal structures for the SpaD backbone pilin, two of which capture the N-terminal domain in either the closed (linear) or open (bent) conformation. To our knowledge, this is the first observation of the bent conformation in Gram-positive pilin structures. Based on this bent conformation, we suggest a three-stage model, which we call the expose-ligate-seal mechanism, for the docking and assembly of backbone pilins into the sortase-dependent pilus.
- Laboratory of Structural Microbiology, Regional Centre for Biotechnology, NCR Biotech Science Cluster, Faridabad, 121001, India.
Organizational Affiliation: