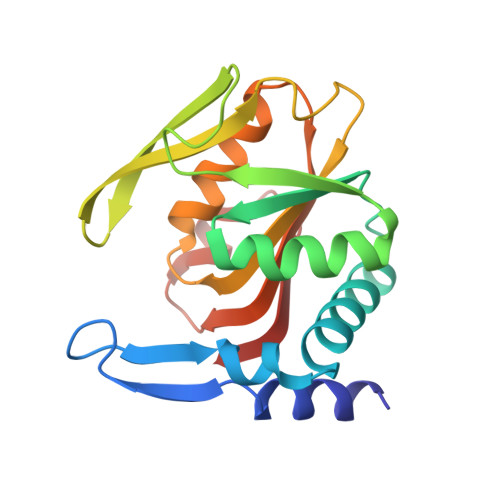
Crystal structures of APRT from Francisella tularensis - an N-H···N hydrogen bond imparts adenine specificity in adenine phosporibosyltransferases.
Pavithra, G.C., Ramagopal, U.A.(2018) FEBS J 285: 2306-2318
- PubMed: 29694705
- DOI: https://doi.org/10.1111/febs.14481
- Primary Citation of Related Structures:
5YW2, 5YW5 - PubMed Abstract:
Francisella tularensisis, the causative agent of tularemia has been classified as a category A bioterrorism agent. Here, we present the crystal structure of apo and adenine bound form of the adenine phosphoribosyltransferase (APRT) from Francisella tularensis. APRT is an enzyme involved in the salvage of adenine (a 6-aminopurine), converting it to AMP. The purine salvage pathway relies on two essential and distinct enzymes to convert 6-aminopurine and 6-oxopurines into corresponding nucleotides. The mechanism by which these enzymes differentiate different purines is not clearly understood. Analysis of the structures of apo and adenine-bound APRT from F. tularensis, together with all other available structures of APRTs, suggests that (a) the base-binding loop is stabilized by a cluster of aromatic and conformation-restricting proline residues, and (b) an N-H···N hydrogen bond between the base-binding loop and the N1 atom of adenine is the key interaction that differentiates adenine from 6-oxopurines. These observations were corroborated by bioinformatics analysis of ~ 4000 sequences of APRTs (with 80% identity cutoff), which confirmed that the residues conferring rigidity to the base-binding loop are highly conserved. Furthermore, an F23A mutation on the base-binding loop severely affects the efficiency of the enzyme. We extended our analysis to the structure and sequences of APRTs from the Trypanosomatidae family with a destabilizing insertion on the base-binding loop and propose the mechanism by which these evolutionarily divergent enzymes achieve base specificity. Our results suggest that the base-binding loop not only confers appropriate affinity but also provides defined specificity for adenine. EC 2.4.2.7 DATABASE: Structural data are available in Protein Data Bank (PDB) under the accession numbers 5YW2 and 5YW5.
- Division of Biological Sciences, Poornaprajna Institute of Scientific Research, Bangalore, India.
Organizational Affiliation: