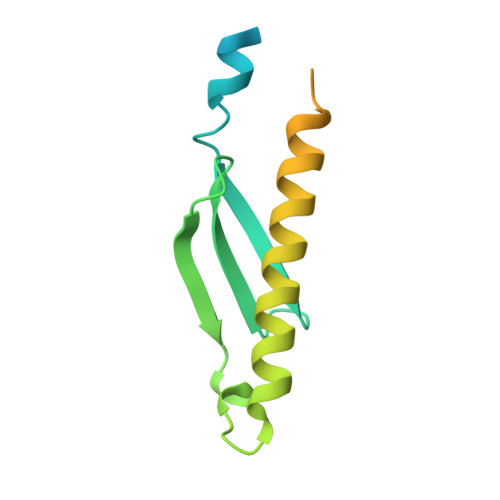
Structural and biophysical characterization of Rv3716c, a hypothetical protein from Mycobacterium tuberculosis
Gopalan, A., Deka, G., Prabhavathi, M., Savithri, H.S., Murthy, M.R.N., Raja, A.(2018) Biochem Biophys Res Commun 495: 982-987
- PubMed: 29154992 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.11.093
- Primary Citation Related Structures:
5YRX - PubMed Abstract:
Latent tuberculosis (TB) is the main hurdle in reaching the goal of "Stop TB 2050". Tuberculin skin and Interferon-gamma release assay tests used currently for the diagnosis of TB infection cannot distinguish between active disease and latent tuberculosis infection (LTBI) and hence new and sensitive protein markers need to be identified for the diagnosis. A protein Rv3716c from Mycobacterium tuberculosis (MtbRv3716c) has been identified as a potential surrogate marker for the diagnosis of LTBI. Here, we present characterization of MtbRv3716c (∼13 kDa) using both biophysical and X-Ray crystallographic methods. EMSA study showed that MtbRv3716c binds to double stranded DNA. X-ray diffraction data collected on a crystal of MtbRv3716c at 1.9 Å resolution was used for structure determination using the molecular replacement method. Significant electron density was not observed for the N-terminal 21 and C-terminal 41 residues in the final electron density map. The C- terminal disordered region is proline rich and displays characteristics of intrinsically disordered proteins. Although the crystal asymmetric unit contained a protomer, a tight dimer could be generated by the application of the crystal two-fold symmetry parallel to the b axis. Packing of dimers in the crystal is mediated by a cadmium ion (Cd 2+ ) occurring at the interface of two dimers. Molecular packing analysis reveals large cavities that are probably occupied by the disordered segments of the N- and C-termini. Structural comparison with other homologous hypothetical DNA binding proteins (PDB codes: 1PUG, 1YBX) highlights structural features that might be significant for DNA binding.
- National Institute for Research in Tuberculosis, Chennai 600031, India.
Organizational Affiliation: