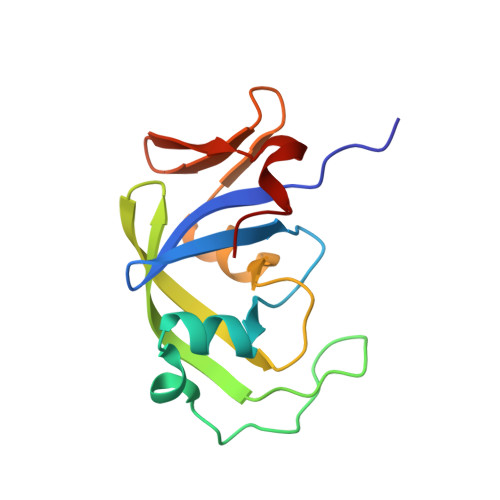
Crystal structure of the retroviral protease-like domain of a protozoal DNA damage-inducible 1 protein.
Kumar, S., Suguna, K.(2018) FEBS Open Bio 8: 1379-1394
- PubMed: 30186740 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/2211-5463.12491
- Primary Citation Related Structures:
5YQ8, 5YS4 - PubMed Abstract:
DNA damage-inducible 1 (Ddi1) is a multidomain protein with one of the domains being retropepsin-like. HIV-1 protease inhibitors were found to reduce opportunistic infections caused by pathogens like Leishmania and Plasmodium , and some of them were shown to inhibit the growth of these parasites. In Leishmania , Ddi1 was identified as a likely target of the inhibitors. We report the crystal structure of the retropepsin-like domain of Ddi1 from Leishmania major as a dimer with clear density for the critical 'flap' region. We have characterized binding with one of the HIV-1 protease inhibitors in solution using bio-layer interferometry and by docking. Further, we have performed molecular dynamics (MD) simulation studies that show that the protein undergoes a conformational change from open to semi-open and closed forms with the closing of the flexible flap over the active site.
- Molecular Biophysics Unit Indian Institute of Science Bangalore India.
Organizational Affiliation: