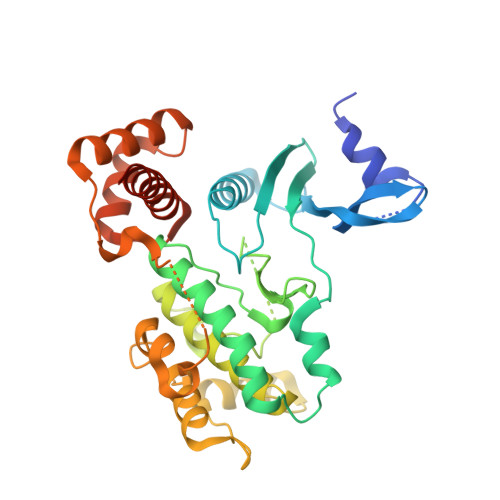
Crystal structure of the kinase and UBA domains of SNRK reveals a distinct UBA binding mode in the AMPK family.
Wang, Y.L., Wang, J., Chen, X., Wang, Z.X., Wu, J.W.(2018) Biochem Biophys Res Commun 495: 1-6
- PubMed: 29061304 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.10.105
- Primary Citation Related Structures:
5YKS - PubMed Abstract:
Sucrose non-fermenting (Snf1)-related kinase (SNRK) is a novel member of the AMP-activated protein kinase (AMPK) family and is involved in many metabolic processes. Here we report the crystal structure of an N-terminal SNRK fragment containing kinase and adjacent ubiquitin-associated (UBA) domains. This structure shows that the UBA domain binds between the N- and C-lobes of the kinase domain. The mode of UBA binding in SNRK largely resembles that in AMPK and brain specific kinase (BRSK), however, unique interactions play vital roles in stabilizing the KD-UBA interface of SNRK. We further propose a potential role of the UBA domain in the regulation of SNRK kinase activity. This study provides new insights into the structural diversities of the AMPK kinase family.
- MOE Key Laboratory of Protein Sciences and Tsinghua-Peking Center for Life Sciences, School of Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: