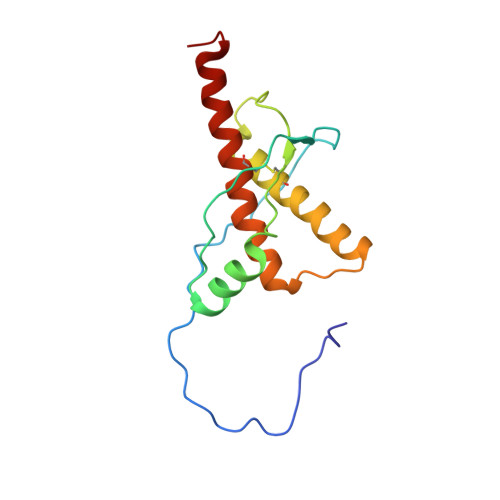
Structural basis for the complete resistance of the human prion protein mutant G127V to prion disease.
Zheng, Z., Zhang, M., Wang, Y., Ma, R., Guo, C., Feng, L., Wu, J., Yao, H., Lin, D.(2018) Sci Rep 8: 13211-13211
- PubMed: 30181558 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-31394-6
- Primary Citation Related Structures:
5YJ4, 5YJ5 - PubMed Abstract:
Prion diseases are caused by the propagation of misfolded cellular prion proteins (PrPs). A completely prion disease-resistant genotype, V127M129, has been identified in Papua New Guinea and verified in transgenic mice. To disclose the structural basis of the disease-resistant effect of the G127V mutant, we determined and compared the structural and dynamic features of the G127V-mutated human PrP (residues 91-231) and the wild-type PrP in solution. HuPrP(G127V) contains α1, α2 and α3 helices and a stretch-strand (SS) pattern comprising residues Tyr128-Gly131 (SS1) and Val161-Arg164 (SS2), with extending atomic distances between the SS1 and SS2 strands, and a structural rearrangement of the Tyr128 side chain due to steric hindrance of the larger hydrophobic side chain of Val127. The extended α1 helix gets closer to the α2 and α3 helices. NMR dynamics analysis revealed that Tyr128, Gly131 and Tyr163 underwent significant conformational exchanges. Molecular dynamics simulations suggest that HuPrP(G127V) prevents the formation of stable β-sheets and dimers. Unique structural and dynamic features potentially inhibit the conformational conversion of the G127V mutant. This work is beneficial for understanding the molecular mechanisms underlying the complete resistance of the G127V mutant to prion disease and for developing new therapeutics for prion disease.
- MOE Key Laboratory of Spectrochemical Analysis & Instrumentation, Key Laboratory of Chemical Biology of Fujian Province, College of Chemistry and Chemical Engineering, Xiamen University, Xiamen, 361005, China.
Organizational Affiliation: