Systematic Profiling of Histone Readers in Arabidopsis thaliana.
Zhao, S., Zhang, B., Yang, M., Zhu, J., Li, H.(2018) Cell Rep 22: 1090-1102
- PubMed: 29386129 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2017.12.099
- Primary Citation Related Structures:
5Y20, 5YC3, 5YC4 - PubMed Abstract:
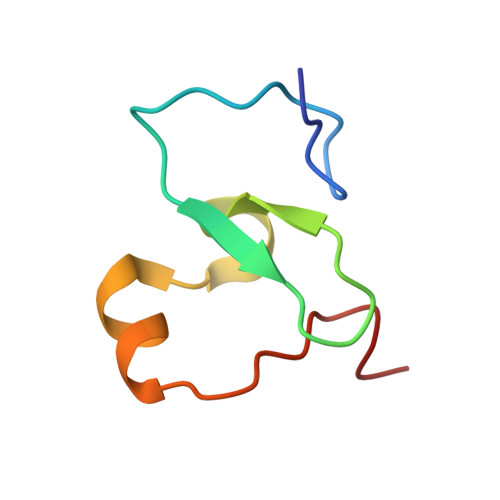
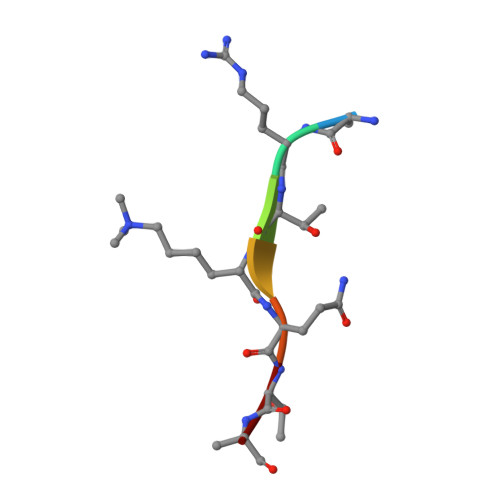
Histone post-translational modifications (PTMs) and their recognition by histone readers exert crucial functions in eukaryotes. Despite extensive studies, conservation and diversity of histone PTM regulation between animals and plants remain less explored because of a lack of systematic knowledge of histone readers in plants. Based on a high-throughput surface plasmon resonance imaging (SPRi) platform, we report the lab-on-chip profiling of interactions between 204 putative reader domains and 11 types of histone peptides in Arabidopsis thaliana. Eleven reader hits were then chosen for histone combinatorial readout pattern profiling. Systematic analysis of histone PTM recognition in Arabidopsis thaliana reveals that plant and human histone readers share conservation in domain types and recognition mechanisms. The differences in particular histone mark recognition by transcription regulator EML1 and DNA damage repair factor MSH6 indicate plant-specific histone PTMs function in Arabidopsis thaliana acquired during evolution.
- MOE Key Laboratory of Protein Sciences, Beijing Advanced Innovation Center for Structural Biology, Department of Basic Medical Sciences, School of Medicine, Tsinghua University, Beijing 100084, China; Tsinghua-Peking Joint Center for Life Sciences, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: