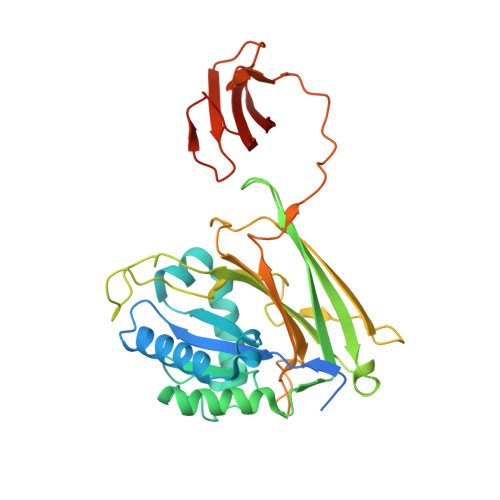
Immunoglobulin-like domains of the cargo proteins are essential for protein stability during secretion by the type IX secretion system.
Sato, K., Kakuda, S., Yukitake, H., Kondo, Y., Shoji, M., Takebe, K., Narita, Y., Naito, M., Nakane, D., Abiko, Y., Hiratsuka, K., Suzuki, M., Nakayama, K.(2018) Mol Microbiol 110: 64-81
- PubMed: 30030863
- DOI: https://doi.org/10.1111/mmi.14083
- Primary Citation of Related Structures:
5Y1A - PubMed Abstract:
The periodontal pathogen Porphyromonas gingivalis secretes many potent virulence factors using the type IX secretion system (T9SS). T9SS cargo proteins that have been structurally determined by X-ray crystallography are composed of a signal peptide, functional domain(s), an immunoglobulin (Ig)-like domain and a C-terminal domain. Role of the Ig-like domains of cargo proteins in the T9SS has not been elucidated. Gingipain proteases, which are cargo proteins of the T9SS, were degraded when their Ig-like domains were lacking or truncated. The degradation was dependent on the activity of a quality control factor, HtrA protease. Another T9SS cargo protein, HBP35, which has a thioredoxin domain as a functional domain, was analyzed by X-ray crystallography, revealing that HBP35 has an Ig-like domain after the thioredoxin domain and that the hydrophobic regions of the thioredoxin domain and the Ig-like domain face each other. HBP35 with substitution of hydrophobic amino acids in the Ig-like domain was degraded depending on HtrA. These results suggest that the Ig-like domain mediates stability of the cargo proteins in the T9SS.
- Department of Microbiology and Oral Infection, Nagasaki University Graduate School of Biomedical Sciences, Nagasaki, Nagasaki, 852-8588, Japan.
Organizational Affiliation: