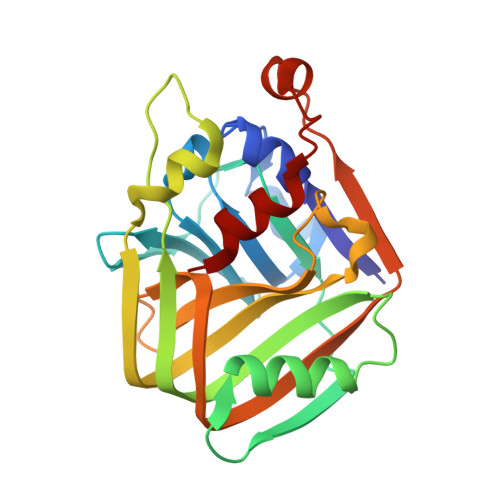
Structural and enzymatic characterization of acetolactate decarboxylase from Bacillus subtilis
Ji, F., Li, M., Feng, Y., Wu, S., Wang, T., Pu, Z., Wang, J., Yang, Y., Xue, S., Bao, Y.(2018) Appl Microbiol Biotechnol 102: 6479-6491
- PubMed: 29796971 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-018-9049-7
- Primary Citation Related Structures:
5XNE - PubMed Abstract:
Acetoin is an important physiological metabolite excreted by microbes. Its functions include avoiding acidification, participating in regulation of the NAD + /NADH ratio, and storing carbon. Acetolactate decarboxylase is a well-characterized anabolic enzyme involved with 3-hydroxy butanone (acetoin). It catalyzes conversion of the (R)- and (S)-enantiomers of acetolactate to generate the single product, (R)-acetoin. In addition to the X-ray crystal structure of acetolactate decarboxylase from Bacillus brevis, although the enzyme is widely present in microorganisms, very few atomic structures of acetolactate decarboxylase are reported. In this paper, we solved and reported a 1.5 Å resolution crystal structure of acetolactate decarboxylase from Bacillus subtilis. Dimeric assembly is observed in the solved structure, which is consistent with the elution profile conducted by molecular filtration. A zinc ion is coordinated by highly conserved histidines (191, 193, and 204) and conserved glutamates (62 and 251). We performed kinetic studies on acetolactate decarboxylase from Bacillus subtilis using circular dichroism, allowing the conversion of acetolactate to chiral acetoin for real-time tracking, yielding a K m value of 21 mM and a k cat value of 2.2 s -1 . Using the two enantiomers of acetolactate as substrates, we further investigated the substrate preference of acetolactate decarboxylase from Bacillus subtilis by means of molecular docking and dynamic simulation in silico. The binding free energy of (S)-acetolactate was found to be ~ 30 kcal/mol greater than that of (R)-acetolactate, indicating a more stable binding for (S)-acetolactate.
- School of Life Science and Biotechnology, Dalian University of Technology, Dalian, Liaoning, 116024, People's Republic of China. fanglingji@dlut.edu.cn.
Organizational Affiliation: