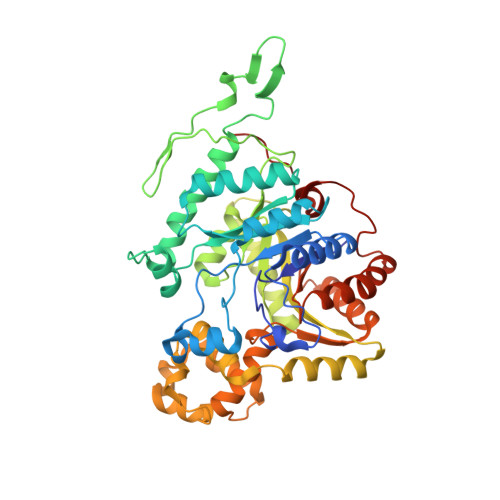
Structural and Biochemical Characterization of BdsA fromBacillus subtilisWU-S2B, a Key Enzyme in the "4S" Desulfurization Pathway.
Su, T., Su, J., Liu, S., Zhang, C., He, J., Huang, Y., Xu, S., Gu, L.(2018) Front Microbiol 9: 231-231
- PubMed: 29497411
- DOI: https://doi.org/10.3389/fmicb.2018.00231
- Primary Citation Related Structures:
5XKC, 5XKD - PubMed Abstract:
Dibenzothiophene (DBT) and their derivatives, accounting for the major part of the sulfur components in crude oil, make one of the most significant pollution sources. The DBT sulfone monooxygenase BdsA, one of the key enzymes in the "4S" desulfurization pathway, catalyzes the oxidation of DBT sulfone to 2'-hydroxybiphenyl 2-sulfonic acid (HBPSi). Here, we determined the crystal structure of BdsA from Bacillus subtilis WU-S2B, at the resolution of 2.2 Å, and the structure of the BdsA-FMN complex at 2.4 Å. BdsA and the BdsA-FMN complex exist as tetramers. DBT sulfone was placed into the active site by molecular docking. Seven residues (Phe12, His20, Phe56, Phe246, Val248, His316, and Val372) are found to be involved in the binding of DBT sulfone. The importance of these residues is supported by the study of the catalytic activity of the active site variants. Structural analysis and enzyme activity assay confirmed the importance of the right position and orientation of FMN and DBT sulfone, as well as the involvement of Ser139 as a nucleophile in catalysis. This work combined with our previous structure of DszC provides a systematic structural basis for the development of engineered desulfurization enzymes with higher efficiency and stability.
- State Key Laboratory of Microbial Technology, School of Life Sciences, Shandong University, Jinan, China.
Organizational Affiliation: