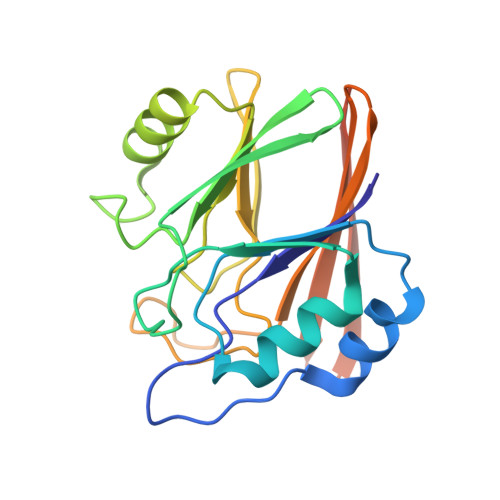
Structural and thermodynamic characterization of metal binding in Vps29 from Entamoeba histolytica: implication in retromer function.
Srivastava, V.K., Yadav, R., Watanabe, N., Tomar, P., Mukherjee, M., Gourinath, S., Nakada-Tsukui, K., Nozaki, T., Datta, S.(2017) Mol Microbiol 106: 562-581
- PubMed: 28898487 Search on PubMed
- DOI: https://doi.org/10.1111/mmi.13836
- Primary Citation Related Structures:
5XCE, 5XCH, 5XCJ, 5XCK - PubMed Abstract:
Vps29 is the smallest subunit of retromer complex with metallo-phosphatase fold. Although the role of metal in Vps29 is in quest, its metal binding mutants has been reported to affect the localization of the retromer complex in human cells. In this study, we report the structural and thermodynamic consequences of these mutations in Vps29 from the protozoan parasite, Entamoeba histolytica (EhVps29). EhVps29 is a zinc binding protein as revealed by X-ray crystallography and isothermal titration calorimetry. The metal binding pocket of EhVps29 exhibits marked differences in its 3-dimensional architecture and metal coordination in comparison to its human homologs and other metallo-phosphatases. Alanine substitutions of the metal-coordinating residues showed significant alteration in the binding affinity of EhVps29 for zinc. We also determined the crystal structures of metal binding defective mutants (D62A and D62A/H86A) of EhVps29. Based on our results, we propose that the metal atoms or the bound water molecules in the metal binding site are important for maintaining the structural integrity of the protein. Further cellular studies in the amoebic trophozoites showed that the overexpression of wild type EhVps29 leads to reduction in intracellular cysteine protease activity suggesting its crucial role in secretion of the proteases.
- Department of Biological Sciences, Indian Institute of Science Education and Research Bhopal, Bhopal 462066, India.
Organizational Affiliation: