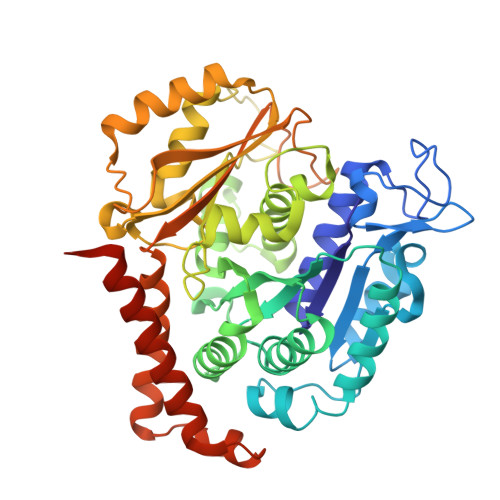
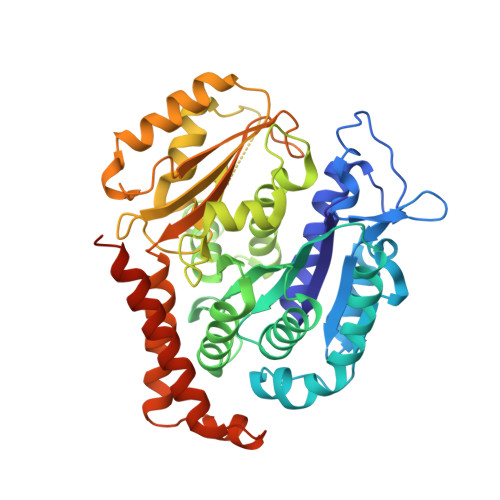
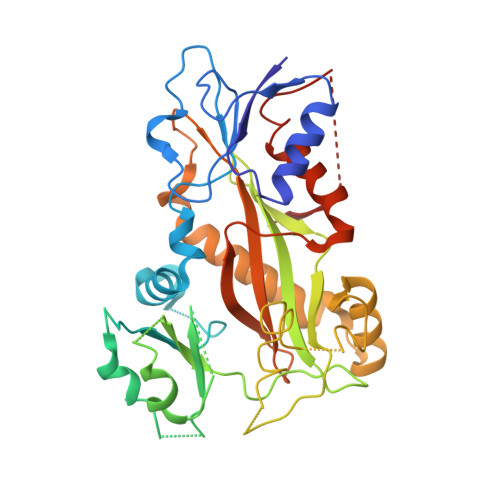
Design, synthesis, biological evaluation and cocrystal structures with tubulin of chiral beta-lactam bridged combretastatin A-4 analogues as potent antitumor agents
Zhou, P., Liang, Y., Zhang, H., Jiang, H., Feng, K., Xu, P., Wang, J., Wang, X., Ding, K., Luo, C., Liu, M., Wang, Y.(2017) Eur J Med Chem 144: 817-842
- PubMed: 29306206 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2017.12.004
- Primary Citation Related Structures:
5XAF, 5XAG - PubMed Abstract:
A diverse of chiral β-lactam bridged analogues of combretastatin A-4 (CA-4), 3-substituted 1,4-diaryl-2-azetidinones, were asymmetrically synthesized and biologically evaluated, leading to identify a number of potent anti-proliferative compounds represented by 14b and 14c with IC 50 values of 0.001-0.021 μM, against four human cancer cell lines (A2780, Hela, SKOV-3 and MDA-MB-231). Structure-activity relationship (SAR) studies on all stereoisomers of 14b and 14c revealed that the absolute configurations of the chiral centers at 3- and 4-position were critically important for the activity and generally a trans configuration between the "A" and "B" rings is optimal. In addition, 14b and 14c displayed less cytotoxicity on normal human oviduct epithelial cells than malignant cells indicating good selectivity in vitro. Further biochemical evaluation and cocrystal structures with tubulin demonstrated that both compounds disrupted tubulin polymerization through interacting at the colchicine-binding site, suppressed angiogenesis in vitro and in vivo, blocked cell cycle progression at mitotic phase and induced cellular apoptosis. The in vivo assays verified that both compounds inhibited xenograft tumor growth in nude mice with acceptable therapeutic window, showing promising potentials for further clinical development.
- School of Pharmacy, Fudan University, Shanghai 201203, China; State Key Laboratory of Organometallic Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai 200032, China.
Organizational Affiliation: