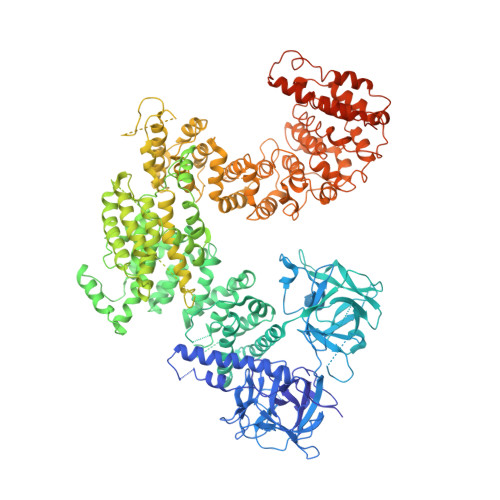
IP3-mediated gating mechanism of the IP3 receptor revealed by mutagenesis and X-ray crystallography
Hamada, K., Miyatake, H., Terauchi, A., Mikoshiba, K.(2017) Proc Natl Acad Sci U S A 114: 4661-4666
- PubMed: 28416699 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1701420114
- Primary Citation Related Structures:
5GUG, 5X9Z, 5XA0, 5XA1 - PubMed Abstract:
The inositol 1,4,5-trisphosphate (IP 3 ) receptor (IP 3 R) is an IP 3 -gated ion channel that releases calcium ions (Ca 2+ ) from the endoplasmic reticulum. The IP 3 -binding sites in the large cytosolic domain are distant from the Ca 2+ conducting pore, and the allosteric mechanism of how IP 3 opens the Ca 2+ channel remains elusive. Here, we identify a long-range gating mechanism uncovered by channel mutagenesis and X-ray crystallography of the large cytosolic domain of mouse type 1 IP 3 R in the absence and presence of IP 3 Analyses of two distinct space group crystals uncovered an IP 3 -dependent global translocation of the curvature α-helical domain interfacing with the cytosolic and channel domains. Mutagenesis of the IP 3 R channel revealed an essential role of a leaflet structure in the α-helical domain. These results suggest that the curvature α-helical domain relays IP 3 -controlled global conformational dynamics to the channel through the leaflet, conferring long-range allosteric coupling from IP 3 binding to the Ca 2+ channel.
- Laboratory for Developmental Neurobiology, Brain Science Institute, RIKEN, Saitama 351-0198, Japan.
Organizational Affiliation: