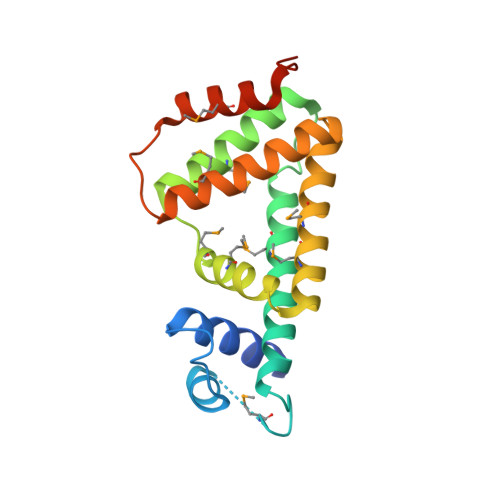
A structural sketch of RcdA, a transcription factor controlling the master regulator of biofilm formation.
Sugino, H., Usui, T., Shimada, T., Nakano, M., Ogasawara, H., Ishihama, A., Hirata, A.(2017) FEBS Lett 591: 2019-2031
- PubMed: 28608551 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.12713
- Primary Citation Related Structures:
5X5I - PubMed Abstract:
RcdA is a regulator of curlin subunit gene D, the master regulator of biofilm formation in Escherichia coli. Here, we determined the X-ray structure of RcdA at 2.55 Å resolution. RcdA consists of an N-terminal DNA-binding domain (DBD) containing a helix-turn-helix (HTH) motif and a C-terminal dimerization domain, and forms a homodimer in crystals. A computational docking model of the RcdA-DNA complex allowed prediction of the candidate residues responsible for DNA binding. Our structure-guided mutagenesis, in combination with gel shift assay, atomic force microscopic observation, and reporter assay, indicate that R32 in α2 of the HTH motif plays an essential role in the recognition and binding of target DNA while T46 in α3 influences the mode of oligomerization. These results provide insights into the DNA-binding mode of RcdA.
- Department of Materials Science and Biotechnology, Graduate School of Science and Engineering, Ehime University, Matsuyama, Japan.
Organizational Affiliation: