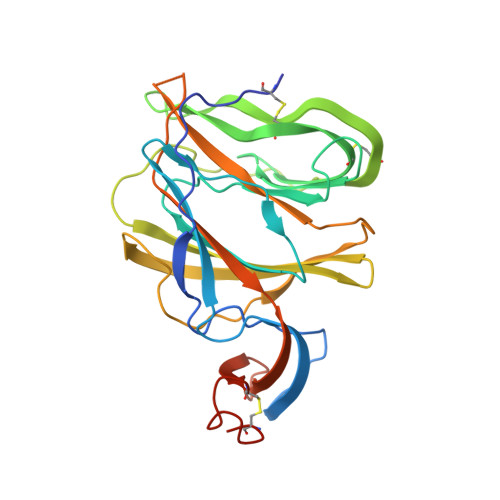
Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains
Yuan, Y., Cao, D., Zhang, Y., Ma, J., Qi, J., Wang, Q., Lu, G., Wu, Y., Yan, J., Shi, Y., Zhang, X., Gao, G.F.(2017) Nat Commun 8: 15092-15092
- PubMed: 28393837 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms15092
- Primary Citation Related Structures:
5X4R, 5X4S, 5X58, 5X59, 5X5B, 5X5C, 5X5F - PubMed Abstract:
The envelope spike (S) proteins of MERS-CoV and SARS-CoV determine the virus host tropism and entry into host cells, and constitute a promising target for the development of prophylactics and therapeutics. Here, we present high-resolution structures of the trimeric MERS-CoV and SARS-CoV S proteins in its pre-fusion conformation by single particle cryo-electron microscopy. The overall structures resemble that from other coronaviruses including HKU1, MHV and NL63 reported recently, with the exception of the receptor binding domain (RBD). We captured two states of the RBD with receptor binding region either buried (lying state) or exposed (standing state), demonstrating an inherently flexible RBD readily recognized by the receptor. Further sequence conservation analysis of six human-infecting coronaviruses revealed that the fusion peptide, HR1 region and the central helix are potential targets for eliciting broadly neutralizing antibodies.
- School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230026, China.
Organizational Affiliation: