Biochemical and structural characterization of Marinomonas mediterranead-mannose isomerase Marme_2490 phylogenetically distant from known enzymes
Saburi, W., Jaito, N., Kato, K., Tanaka, Y., Yao, M., Mori, H.(2018) Biochimie 144: 63-73
- PubMed: 29107017 Search on PubMed
- DOI: https://doi.org/10.1016/j.biochi.2017.10.016
- Primary Citation Related Structures:
5X32 - PubMed Abstract:
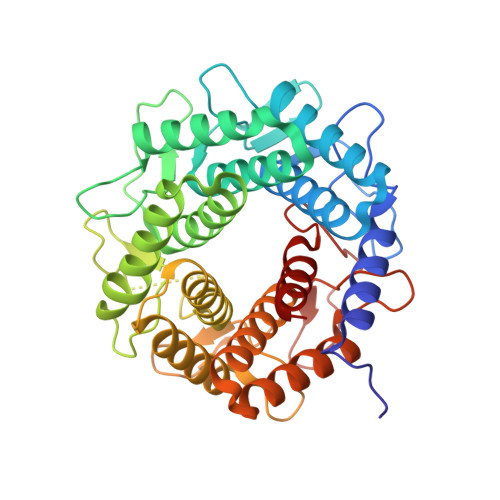
d-Mannose isomerase (MI) reversibly isomerizes d-mannose to d-fructose, and is attractive for producing d-mannose from inexpensive d-fructose. It belongs to the N-acylglucosamine 2-epimerase (AGE) superfamily along with AGE, cellobiose 2-epimerase (CE), and aldose-ketose isomerase (AKI). In this study, Marinomonas mediterranea Marme_2490, showing low sequence identity with any known enzymes, was found to isomerize d-mannose as its primary substrate. Marme_2490 also isomerized d-lyxose and 4-OH d-mannose derivatives (d-talose and 4-O-monosaccharyl-d-mannose). Its activity for d-lyxose is known in other d-mannose isomerizing enzymes, such as MI and AKI, but we identified, for the first time, its activity for 4-OH d-mannose derivatives. Marme_2490 did not isomerize d-glucose, as known MIs do not, while AKI isomerizes both d-mannose and d-glucose. Thus, Marme_2490 was concluded to be an MI. The initial and equilibrium reaction products were analyzed by NMR to illuminate mechanistic information regarding the Marme_2490 reaction. The analysis of the initial reaction product revealed that β-d-mannose was formed. In the analysis of the equilibrated reaction products in D 2 O, signals of 2-H of d-mannose and 1-H of d-fructose were clearly detected. This indicates that these protons are not substituted with deuterium from D 2 O and Marme_2490 catalyzes the intramolecular proton transfer between 1-C and 2-C. The crystal structure of Marme_2490 in a ligand-free form was determined and found that Marme_2490 is formed by an (α/α) 6 -barrel, which is commonly observed in AGE superfamily enzymes. Despite diverse reaction specificities, the orientations of residues involved in catalysis and substrate binding by Marme_2490 were similar to those in both AKI (Salmonella enterica AKI) and epimerase (Rhodothermus marinus CE). The Marme_2490 structure suggested that the α7→α8 and α11→α12 loops of the catalytic domain participated in the formation of an open substrate-binding site to provide sufficient space to bind 4-OH d-mannose derivatives.
- Research Faculty of Agriculture, Hokkaido University, N-9, W-9, Sapporo 060-8589, Japan.
Organizational Affiliation: