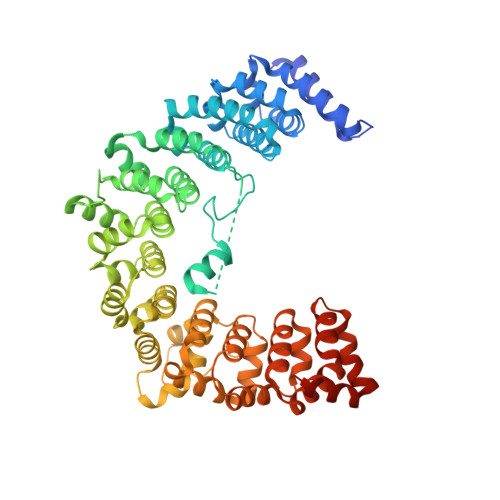
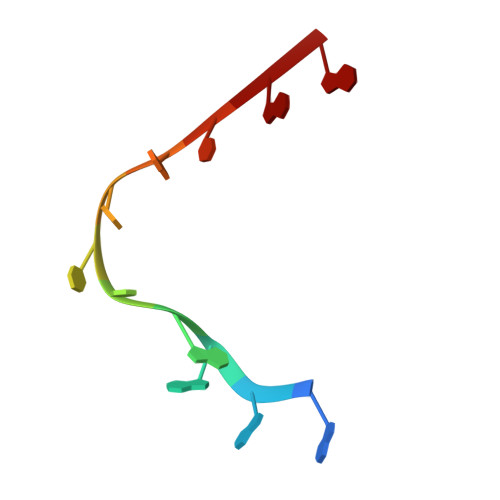
Structural basis for the specific recognition of 18S rRNA by APUM23.
Bao, H., Wang, N., Wang, C., Jiang, Y., Liu, J., Xu, L., Wu, J., Shi, Y.(2017) Nucleic Acids Res 45: 12005-12014
- PubMed: 29036323
- DOI: https://doi.org/10.1093/nar/gkx872
- Primary Citation of Related Structures:
5WZG, 5WZH, 5WZI, 5WZJ, 5WZK - PubMed Abstract:
PUF (Pumilio/fem-3 mRNA binding factor) proteins, a conserved family of RNA-binding proteins, recognize specific single-strand RNA targets in a specific modular way. Although plants have a greater number of PUF protein members than do animal and fungal systems, they have been the subject of fewer structural and functional investigations. The aim of this study was to elucidate the involvement of APUM23, a nucleolar PUF protein in the plant Arabidopsis, in pre-rRNA processing. APUM23 is distinct from classical PUF family proteins, which are located in the cytoplasm and bind to 3'UTRs of mRNA to modulate mRNA expression and localization. We found that the complete RNA target sequence of APUM23 comprises 11 nt in 18S rRNA at positions 1141-1151. The complex structure shows that APUM23 has 10 PUF repeats; it assembles into a C-shape, with an insertion located within the inner concave surface. We found several different RNA recognition features. A notable structural feature of APUM23 is an insertion in the third PUF repeat that participates in nucleotide recognition and maintains the correct conformation of the target RNA. Our findings elucidate the mechanism for APUM23's-specific recognition of 18S rRNA.
- Hefei National Laboratory for Physical Science at Microscale, School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230027, China.
Organizational Affiliation: