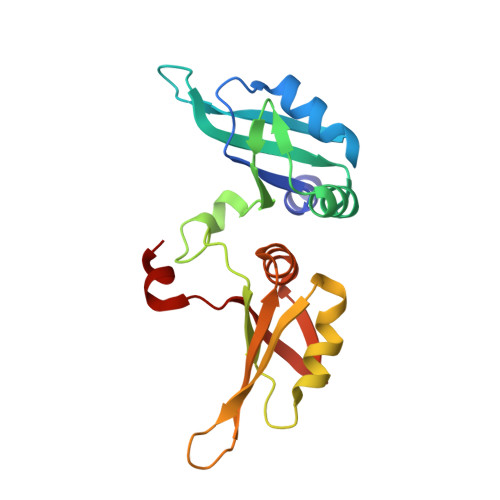
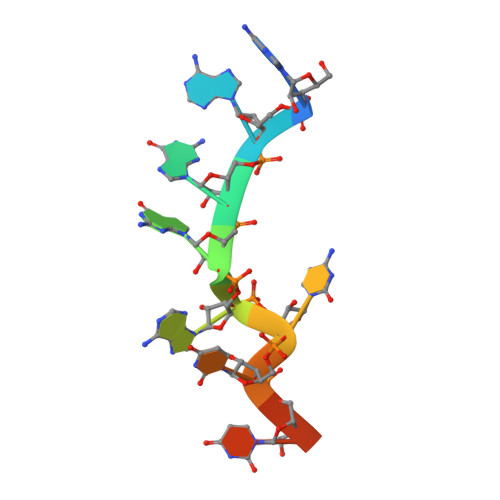
Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1.
Wu, B., Su, S., Patil, D.P., Liu, H., Gan, J., Jaffrey, S.R., Ma, J.(2018) Nat Commun 9: 420-420
- PubMed: 29379020 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02770-z
- Primary Citation Related Structures:
5HO4, 5WWE, 5WWF, 5WWG - PubMed Abstract:
Human hnRNP A2/B1 is an RNA-binding protein that plays important roles in many biological processes, including maturation, transport, and metabolism of mRNA, and gene regulation of long noncoding RNAs. hnRNP A2/B1 was reported to control the microRNAs sorting to exosomes and promote primary microRNA processing as a potential m 6 A "reader." hnRNP A2/B1 contains two RNA recognition motifs that provide sequence-specific recognition of RNA substrates. Here, we determine crystal structures of tandem RRM domains of hnRNP A2/B1 in complex with various RNA substrates, elucidating specific recognitions of AGG and UAG motifs by RRM1 and RRM2 domains, respectively. Further structural and biochemical results demonstrate multivariant binding modes for sequence-diversified RNA substrates, supporting a RNA matchmaker mechanism in hnRNP A2/B1 function. Moreover, our studies in combination with bioinformatic analysis suggest that hnRNP A2/B1 may mediate effects of m 6 A through a "m 6 A switch" mechanism, instead of acting as a direct "reader" of m 6 A modification.
- State Key Laboratory of Genetic Engineering, Collaborative Innovation Centre of Genetics and Development, Department of Biochemistry, Institute of Plant Biology, School of Life Sciences, Fudan University, Shanghai, 200438, China.
Organizational Affiliation: