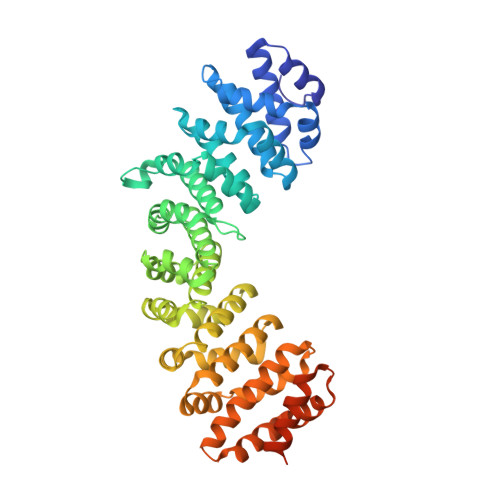
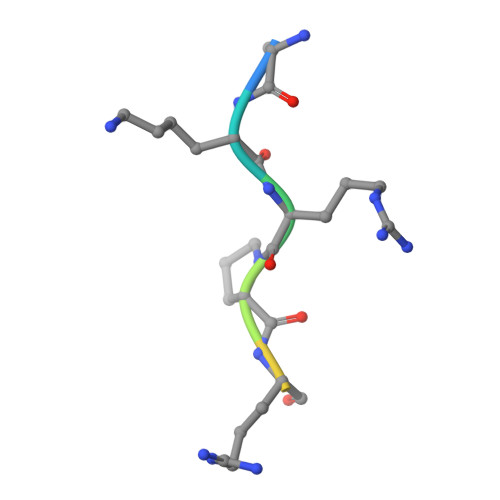
Structural basis for the regulation of nuclear import of Epstein-Barr virus nuclear antigen 1 (EBNA1) by phosphorylation of the nuclear localization signal.
Nakada, R., Hirano, H., Matsuura, Y.(2017) Biochem Biophys Res Commun 484: 113-117
- PubMed: 28104399 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2017.01.063
- Primary Citation Related Structures:
5WUM, 5WUN - PubMed Abstract:
Epstein-Barr virus (EBV) nuclear antigen 1 (EBNA1) is expressed in every EBV-positive tumor and is essential for the maintenance, replication, and transcription of the EBV genome in the nucleus of host cells. EBNA1 is a serine phosphoprotein, and it has been shown that phosphorylation of S385 in the nuclear localization signal (NLS) of EBNA1 increases the binding affinity to the nuclear import adaptor importin-α1 as well as importin-α5, and stimulates nuclear import of EBNA1. To gain insights into how phosphorylation of the EBNA1 NLS regulates nuclear import, we have determined the crystal structures of two peptide complexes of importin-α1: one with S385-phosphorylated EBNA1 NLS peptide, determined at 2.0 Å resolution, and one with non-phosphorylated EBNA1 NLS peptide, determined at 2.2 Å resolution. The structures show that EBNA1 NLS binds to the major and minor NLS-binding sites of importin-α1, and indicate that the binding affinity of the EBNA1 NLS to the minor NLS-binding site could be enhanced by phosphorylation of S385 through electrostatic interaction between the phosphate group of phospho-S385 and K392 of importin-α1 (corresponding to R395 of importin-α5) on armadillo repeat 8.
- Division of Biological Science, Graduate School of Science, Nagoya University, Japan.
Organizational Affiliation: